TLR-9 and IL-15 Synergy Promotes the In Vitro Clonal Expansion of Chronic Lymphocytic Leukemia B Cells
- PMID: 26136429
- PMCID: PMC4505957
- DOI: 10.4049/jimmunol.1403189
TLR-9 and IL-15 Synergy Promotes the In Vitro Clonal Expansion of Chronic Lymphocytic Leukemia B Cells
Abstract
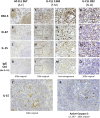
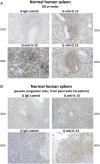
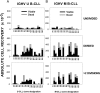
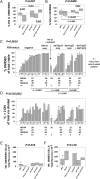
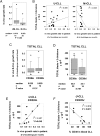
Clinical progression of B cell chronic lymphocytic leukemia (B-CLL) reflects the clone's Ag receptor (BCR) and involves stroma-dependent B-CLL growth within lymphoid tissue. Uniformly elevated expression of TLR-9, occasional MYD88 mutations, and BCR specificity for DNA or Ags physically linked to DNA together suggest that TLR-9 signaling is important in driving B-CLL growth in patients. Nevertheless, reports of apoptosis after B-CLL exposure to CpG oligodeoxynucleotide (ODN) raised questions about a central role for TLR-9. Because normal memory B cells proliferate vigorously to ODN+IL-15, a cytokine found in stromal cells of bone marrow, lymph nodes, and spleen, we examined whether this was true for B-CLL cells. Through a CFSE-based assay for quantitatively monitoring in vitro clonal proliferation/survival, we show that IL-15 precludes TLR-9-induced apoptosis and permits significant B-CLL clonal expansion regardless of the clone's BCR mutation status. A robust response to ODN+IL-15 was positively linked to presence of chromosomal anomalies (trisomy-12 or ataxia telangiectasia mutated anomaly + del13q14) and negatively linked to a very high proportion of CD38(+) cells within the blood-derived B-CLL population. Furthermore, a clone's intrinsic potential for in vitro growth correlated directly with doubling time in blood, in the case of B-CLL with Ig H chain V region-unmutated BCR and <30% CD38(+) cells in blood. Finally, in vitro high-proliferator status was statistically linked to diminished patient survival. These findings, together with immunohistochemical evidence of apoptotic cells and IL-15-producing cells proximal to B-CLL pseudofollicles in patient spleens, suggest that collaborative ODN and IL-15 signaling may promote in vivo B-CLL growth.
Copyright © 2015 by The American Association of Immunologists, Inc.
Figures






Similar articles
-
Mechanistic Insights into CpG DNA and IL-15 Synergy in Promoting B Cell Chronic Lymphocytic Leukemia Clonal Expansion.J Immunol. 2018 Sep 1;201(5):1570-1585. doi: 10.4049/jimmunol.1800591. Epub 2018 Aug 1. J Immunol. 2018. PMID: 30068596 Free PMC article.
-
Heterogeneous functional effects of concomitant B cell receptor and TLR stimulation in chronic lymphocytic leukemia with mutated versus unmutated Ig genes.J Immunol. 2014 May 15;192(10):4518-24. doi: 10.4049/jimmunol.1302102. Epub 2014 Apr 9. J Immunol. 2014. PMID: 24719462 Clinical Trial.
-
Mechanism for IL-15-Driven B Cell Chronic Lymphocytic Leukemia Cycling: Roles for AKT and STAT5 in Modulating Cyclin D2 and DNA Damage Response Proteins.J Immunol. 2019 May 15;202(10):2924-2944. doi: 10.4049/jimmunol.1801142. Epub 2019 Apr 15. J Immunol. 2019. PMID: 30988120 Free PMC article.
-
[Research progress on prognostic markers of chronic lymphocytic leukemia].Zhejiang Da Xue Xue Bao Yi Xue Ban. 2010 May;39(3):250-6. doi: 10.3785/j.issn.1008-9292.2010.03.007. Zhejiang Da Xue Xue Bao Yi Xue Ban. 2010. PMID: 20544986 Review. Chinese.
-
Stereotyped B-cell receptors in chronic lymphocytic leukemia.Leuk Lymphoma. 2014 Oct;55(10):2252-61. doi: 10.3109/10428194.2013.879715. Epub 2014 Mar 17. Leuk Lymphoma. 2014. PMID: 24397617 Review.
Cited by
-
The Role of the Microenvironment and Cell Adhesion Molecules in Chronic Lymphocytic Leukemia.Cancers (Basel). 2023 Oct 26;15(21):5160. doi: 10.3390/cancers15215160. Cancers (Basel). 2023. PMID: 37958334 Free PMC article. Review.
-
Toll-like receptor 9 signaling in chronic lymphocytic leukemia cell lines.FEBS Open Bio. 2023 Dec;13(12):2367-2374. doi: 10.1002/2211-5463.13726. Epub 2023 Nov 8. FEBS Open Bio. 2023. PMID: 37881888 Free PMC article.
-
Extrinsic interactions in the microenvironment in vivo activate an antiapoptotic multidrug-resistant phenotype in CLL.Blood Adv. 2021 Sep 14;5(17):3497-3510. doi: 10.1182/bloodadvances.2020003944. Blood Adv. 2021. PMID: 34432864 Free PMC article.
-
Characterization of the Intraclonal Complexity of Chronic Lymphocytic Leukemia B Cells: Potential Influences of B-Cell Receptor Crosstalk with Other Stimuli.Cancers (Basel). 2023 Sep 25;15(19):4706. doi: 10.3390/cancers15194706. Cancers (Basel). 2023. PMID: 37835400 Free PMC article.
-
Ibrutinib sensitizes CLL cells to venetoclax by interrupting TLR9-induced CD40 upregulation and protein translation.Leukemia. 2023 Jun;37(6):1268-1276. doi: 10.1038/s41375-023-01898-w. Epub 2023 Apr 26. Leukemia. 2023. PMID: 37100883 Free PMC article.
References
-
- Redaelli A., Laskin B. L., Stephens J. M., Botteman M. F., Pashos C. L. 2004. The clinical and epidemiological burden of chronic lymphocytic leukaemia. Eur. J. Cancer Care (Engl.) 13: 279–287. - PubMed
-
- van Gent R., Kater A. P., Otto S. A., Jaspers A., Borghans J. A., Vrisekoop N., Ackermans M. A., Ruiter A. F., Wittebol S., Eldering E., et al. 2008. In vivo dynamics of stable chronic lymphocytic leukemia inversely correlate with somatic hypermutation levels and suggest no major leukemic turnover in bone marrow. Cancer Res. 68: 10137–10144. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

