Systems biology and gene networks in neurodevelopmental and neurodegenerative disorders
- PMID: 26149713
- PMCID: PMC4699316
- DOI: 10.1038/nrg3934
Systems biology and gene networks in neurodevelopmental and neurodegenerative disorders
Abstract
Genetic and genomic approaches have implicated hundreds of genetic loci in neurodevelopmental disorders and neurodegeneration, but mechanistic understanding continues to lag behind the pace of gene discovery. Understanding the role of specific genetic variants in the brain involves dissecting a functional hierarchy that encompasses molecular pathways, diverse cell types, neural circuits and, ultimately, cognition and behaviour. With a focus on transcriptomics, this Review discusses how high-throughput molecular, integrative and network approaches inform disease biology by placing human genetics in a molecular systems and neurobiological context. We provide a framework for interpreting network biology studies and leveraging big genomics data sets in neurobiology.
Conflict of interest statement
The authors declare no competing interests.
Figures


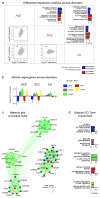
References
-
- Gratten J, Wray NR, Keller MC, Visscher PM. Large-scale genomics unveils the genetic architecture of psychiatric disorders. Nat Neurosci. 2014;17:782–790. A comprehensive review of GWASs and exome studies across major neuropsychiatric disorders. It discusses the role of common variants and rare variants across disorders, and the concept of explaining heritability. - PMC - PubMed
-
- Bullmore E, Sporns O. Complex brain networks: graph theoretical analysis of structural and functional systems. Nat Rev Neurosci. 2009;10:186–198. - PubMed
-
- Grant S. Systems biology in neuroscience: bridging genes to cognition. Curr Opin Neurobiol. 2003;13:577–582. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

