Synthetic biosensors for precise gene control and real-time monitoring of metabolites
- PMID: 26152303
- PMCID: PMC4551912
- DOI: 10.1093/nar/gkv616
Synthetic biosensors for precise gene control and real-time monitoring of metabolites
Abstract
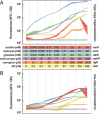
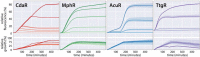
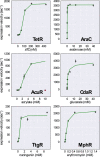
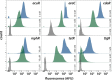
Characterization and standardization of inducible transcriptional regulators has transformed how scientists approach biology by allowing precise and tunable control of gene expression. Despite their utility, only a handful of well-characterized regulators exist, limiting the complexity of engineered biological systems. We apply a characterization pipeline to four genetically encoded sensors that respond to acrylate, glucarate, erythromycin and naringenin. We evaluate how the concentration of the inducing chemical relates to protein expression, how the extent of induction affects protein expression kinetics, and how the activation behavior of single cells relates to ensemble measurements. We show that activation of each sensor is orthogonal to the other sensors, and to other common inducible systems. We demonstrate independent control of three fluorescent proteins in a single cell, chemically defining eight unique transcriptional states. To demonstrate biosensor utility in metabolic engineering, we apply the glucarate biosensor to monitor product formation in a heterologous glucarate biosynthesis pathway and identify superior enzyme variants. Doubling the number of well-characterized inducible systems makes more complex synthetic biological circuits accessible. Characterizing sensors that transduce the intracellular concentration of valuable metabolites into fluorescent readouts enables high-throughput screening of biological catalysts and alleviates the primary bottleneck of the metabolic engineering design-build-test cycle.
© The Author(s) 2015. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

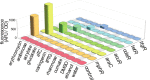
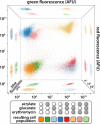
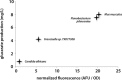
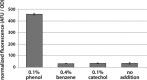
Similar articles
-
Design and application of genetically-encoded malonyl-CoA biosensors for metabolic engineering of microbial cell factories.Metab Eng. 2017 Nov;44:253-264. doi: 10.1016/j.ymben.2017.10.011. Epub 2017 Oct 31. Metab Eng. 2017. PMID: 29097310 Review.
-
Design and Characterization of Biosensors for the Screening of Modular Assembled Naringenin Biosynthetic Library in Saccharomyces cerevisiae.ACS Synth Biol. 2019 Sep 20;8(9):2121-2130. doi: 10.1021/acssynbio.9b00212. Epub 2019 Aug 28. ACS Synth Biol. 2019. PMID: 31433622
-
Genetically encoded sensors enable real-time observation of metabolite production.Proc Natl Acad Sci U S A. 2016 Mar 1;113(9):2388-93. doi: 10.1073/pnas.1600375113. Epub 2016 Feb 8. Proc Natl Acad Sci U S A. 2016. PMID: 26858408 Free PMC article.
-
Microbial Transcription Factor-Based Biosensors: Innovations from Design to Applications in Synthetic Biology.Biosensors (Basel). 2025 Mar 31;15(4):221. doi: 10.3390/bios15040221. Biosensors (Basel). 2025. PMID: 40277535 Free PMC article. Review.
-
Applications of genetically-encoded biosensors for the construction and control of biosynthetic pathways.Metab Eng. 2012 May;14(3):212-22. doi: 10.1016/j.ymben.2011.09.004. Epub 2011 Sep 18. Metab Eng. 2012. PMID: 21946159 Free PMC article. Review.
Cited by
-
Cell factories for biosynthesis of D-glucaric acid: a fusion of static and dynamic strategies.World J Microbiol Biotechnol. 2024 Aug 8;40(10):292. doi: 10.1007/s11274-024-04097-6. World J Microbiol Biotechnol. 2024. PMID: 39112688 Review.
-
Metabolically-targeted dCas9 expression in bacteria.Nucleic Acids Res. 2023 Jan 25;51(2):982-996. doi: 10.1093/nar/gkac1248. Nucleic Acids Res. 2023. PMID: 36629257 Free PMC article.
-
In vivo biosensors: mechanisms, development, and applications.J Ind Microbiol Biotechnol. 2018 Jul;45(7):491-516. doi: 10.1007/s10295-018-2004-x. Epub 2018 Jan 29. J Ind Microbiol Biotechnol. 2018. PMID: 29380152
-
Development of novel metabolite-responsive transcription factors via transposon-mediated protein fusion.Protein Eng Des Sel. 2018 Feb 1;31(2):55-63. doi: 10.1093/protein/gzy001. Protein Eng Des Sel. 2018. PMID: 29385546 Free PMC article.
-
Tetramethylpyrazine-Inducible Promoter Region from Rhodococcus jostii TMP1.Molecules. 2018 Jun 25;23(7):1530. doi: 10.3390/molecules23071530. Molecules. 2018. PMID: 29941849 Free PMC article.
References
-
- Baneyx F. Recombinant protein expression in Escherichia coli. Curr. Opin. Biotechnol. 1999;10:411–421. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

