Entry of Francisella tularensis into Murine B Cells: The Role of B Cell Receptors and Complement Receptors
- PMID: 26161475
- PMCID: PMC4498600
- DOI: 10.1371/journal.pone.0132571
Entry of Francisella tularensis into Murine B Cells: The Role of B Cell Receptors and Complement Receptors
Abstract
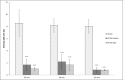
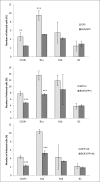
Francisella tularensis, the etiological agent of tularemia, is an intracellular pathogen that dominantly infects and proliferates inside phagocytic cells but can be seen also in non-phagocytic cells, including B cells. Although protective immunity is known to be almost exclusively associated with the type 1 pathway of cellular immunity, a significant role of B cells in immune responses already has been demonstrated. Whether their role is associated with antibody-dependent or antibody-independent B cell functions is not yet fully understood. The character of early events during B cell-pathogen interaction may determine the type of B cell response regulating the induction of adaptive immunity. We used fluorescence microscopy and flow cytometry to identify the basic requirements for the entry of F. tularensis into B cells within in vivo and in vitro infection models. Here, we present data showing that Francisella tularensis subsp. holarctica strain LVS significantly infects individual subsets of murine peritoneal B cells early after infection. Depending on a given B cell subset, uptake of Francisella into B cells is mediated by B cell receptors (BCRs) with or without complement receptor CR1/2. However, F. tularensis strain FSC200 ΔiglC and ΔftdsbA deletion mutants are defective in the ability to enter B cells. Once internalized into B cells, F. tularensis LVS intracellular trafficking occurs along the endosomal pathway, albeit without significant multiplication. The results strongly suggest that BCRs alone within the B-1a subset can ensure the internalization process while the BCRs on B-1b and B-2 cells need co-signaling from the co receptor containing CR1/2 to initiate F. tularensis engulfment. In this case, fluidity of the surface cell membrane is a prerequisite for the bacteria's internalization. The results substantially underline the functional heterogeneity of B cell subsets in relation to F. tularensis.
Conflict of interest statement
Figures




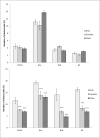
Similar articles
-
Innate Immune Recognition: Implications for the Interaction of Francisella tularensis with the Host Immune System.Front Cell Infect Microbiol. 2017 Oct 16;7:446. doi: 10.3389/fcimb.2017.00446. eCollection 2017. Front Cell Infect Microbiol. 2017. PMID: 29085810 Free PMC article. Review.
-
A novel receptor - ligand pathway for entry of Francisella tularensis in monocyte-like THP-1 cells: interaction between surface nucleolin and bacterial elongation factor Tu.BMC Microbiol. 2008 Sep 12;8:145. doi: 10.1186/1471-2180-8-145. BMC Microbiol. 2008. PMID: 18789156 Free PMC article.
-
Deletion of IglH in virulent Francisella tularensis subsp. holarctica FSC200 strain results in attenuation and provides protection against the challenge with the parental strain.Microbes Infect. 2012 Feb;14(2):177-87. doi: 10.1016/j.micinf.2011.08.017. Epub 2011 Sep 8. Microbes Infect. 2012. PMID: 21930232
-
B cell subsets are activated and produce cytokines during early phases of Francisella tularensis LVS infection.Microb Pathog. 2014 Oct;75:49-58. doi: 10.1016/j.micpath.2014.08.009. Epub 2014 Sep 6. Microb Pathog. 2014. PMID: 25200734
-
Francisella tularensis Susceptibility to Antibiotics: A Comprehensive Review of the Data Obtained In vitro and in Animal Models.Front Cell Infect Microbiol. 2017 Apr 11;7:122. doi: 10.3389/fcimb.2017.00122. eCollection 2017. Front Cell Infect Microbiol. 2017. PMID: 28443249 Free PMC article. Review.
Cited by
-
Francisella tularensis Confronts the Complement System.Front Cell Infect Microbiol. 2017 Dec 19;7:523. doi: 10.3389/fcimb.2017.00523. eCollection 2017. Front Cell Infect Microbiol. 2017. PMID: 29312899 Free PMC article.
-
Innate Immune Recognition: Implications for the Interaction of Francisella tularensis with the Host Immune System.Front Cell Infect Microbiol. 2017 Oct 16;7:446. doi: 10.3389/fcimb.2017.00446. eCollection 2017. Front Cell Infect Microbiol. 2017. PMID: 29085810 Free PMC article. Review.
-
Why Does SARS-CoV-2 Infection Induce Autoantibody Production?Pathogens. 2021 Mar 22;10(3):380. doi: 10.3390/pathogens10030380. Pathogens. 2021. PMID: 33809954 Free PMC article.
-
Autotransporter-Mediated Display of Complement Receptor Ligands by Gram-Negative Bacteria Increases Antibody Responses and Limits Disease Severity.Pathogens. 2020 May 14;9(5):375. doi: 10.3390/pathogens9050375. Pathogens. 2020. PMID: 32422907 Free PMC article.
-
Innate Immune Recognition: An Issue More Complex Than Expected.Front Cell Infect Microbiol. 2019 Jul 3;9:241. doi: 10.3389/fcimb.2019.00241. eCollection 2019. Front Cell Infect Microbiol. 2019. PMID: 31334134 Free PMC article. Review.
References
-
- Pizarro-Cerdá J, Moreno E, Gorvel JP. Invasion and intracellular trafficking of Brucella abortus in nonphagocytic cells. Microbes Infect Inst Pasteur. 2000;2: 829–835. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

