A Crystallin Fold in the Interleukin-4-inducing Principle of Schistosoma mansoni Eggs (IPSE/α-1) Mediates IgE Binding for Antigen-independent Basophil Activation
- PMID: 26163514
- PMCID: PMC4571962
- DOI: 10.1074/jbc.M115.675066
A Crystallin Fold in the Interleukin-4-inducing Principle of Schistosoma mansoni Eggs (IPSE/α-1) Mediates IgE Binding for Antigen-independent Basophil Activation
Abstract
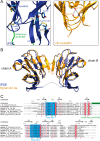
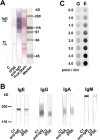
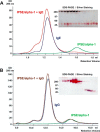
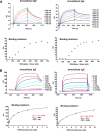
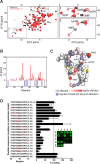
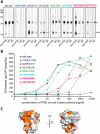
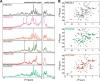
The IL-4-inducing principle from Schistosoma mansoni eggs (IPSE/α-1), the major secretory product of eggs from the parasitic worm S. mansoni, efficiently triggers basophils to release the immunomodulatory key cytokine interleukin-4. Activation by IPSE/α-1 requires the presence of IgE on the basophils, but the detailed molecular mechanism underlying activation is unknown. NMR and crystallographic analysis of IPSEΔNLS, a monomeric IPSE/α-1 mutant, revealed that IPSE/α-1 is a new member of the βγ-crystallin superfamily. We demonstrate that this molecule is a general immunoglobulin-binding factor with highest affinity for IgE. NMR binding studies of IPSEΔNLS with the 180-kDa molecule IgE identified a large positively charged binding surface that includes a flexible loop, which is unique to the IPSE/α-1 crystallin fold. Mutational analysis of amino acids in the binding interface showed that residues contributing to IgE binding are important for IgE-dependent activation of basophils. As IPSE/α-1 is unable to cross-link IgE, we propose that this molecule, by taking advantage of its unique IgE-binding crystallin fold, activates basophils by a novel, cross-linking-independent mechanism.
Keywords: Schistosoma mansoni; basophil; crystal structure; crystallin; interleukin; nuclear magnetic resonance (NMR).
© 2015 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures




References
-
- Schramm G., Haas H. (2010) Th2 immune response against Schistosoma mansoni infection. Microbes Infect. 12, 881–888 - PubMed
-
- Brunet L. R., Finkelman F. D., Cheever A. W., Kopf M. A., Pearce E. J. (1997) IL-4 protects against TNF-α 32 -mediated cachexia and death during acute schistosomiasis. J. Immunol. 159, 777–785 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Research Materials

