New classes of self-cleaving ribozymes revealed by comparative genomics analysis
- PMID: 26167874
- PMCID: PMC4509812
- DOI: 10.1038/nchembio.1846
New classes of self-cleaving ribozymes revealed by comparative genomics analysis
Abstract
Enzymes made of RNA catalyze reactions that are essential for protein synthesis and RNA processing. However, such natural ribozymes are exceedingly rare, as evidenced by the fact that the discovery rate for new classes has dropped to one per decade from about one per year during the 1980s. Indeed, only 11 distinct ribozyme classes have been experimentally validated to date. Recently, we recognized that self-cleaving ribozymes frequently associate with certain types of genes from bacteria. Herein we exploited this association to identify divergent architectures for two previously known ribozyme classes and to discover additional noncoding RNA motifs that are self-cleaving RNA candidates. We identified three new self-cleaving classes, which we named twister sister, pistol and hatchet, from this collection, suggesting that even more ribozymes remain hidden in modern cells.
Conflict of interest statement
The authors declare no competing financial interests.
Figures
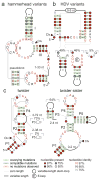


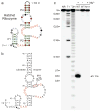
References
-
- Nissen P, Hansen J, Ban N, Moore PB, Steitz TA. The structural basis of ribosome activity in peptide bond synthesis. Science. 2000;289:920–930. - PubMed
-
- Lambowitz AM, Caprara MG, Zimmerly S, Perlman PS. Group I and Group II Ribozymes as RNPs: Clues to the Past and Guides to the Future. In: Gesteland RF, Cech TR, Atkins JF, editors. The RNA World. 2. Vol. 37. Cold Spring Harbor Laboratory Press; Cold Spring Harbor: 1999. pp. 451–483.
-
- Ellis JC, Brown JW. The RNase P family. RNA Biol. 2009;6:362–369. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

