Genome-Wide MicroRNA and Gene Analysis of Mesenchymal Stem Cell Chondrogenesis Identifies an Essential Role and Multiple Targets for miR-140-5p
- PMID: 26175215
- PMCID: PMC4737122
- DOI: 10.1002/stem.2093
Genome-Wide MicroRNA and Gene Analysis of Mesenchymal Stem Cell Chondrogenesis Identifies an Essential Role and Multiple Targets for miR-140-5p
Abstract
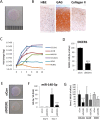
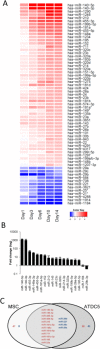
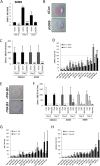
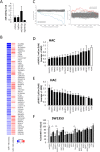
microRNAs (miRNAs) are abundantly expressed in development where they are critical determinants of cell differentiation and phenotype. Accordingly miRNAs are essential for normal skeletal development and chondrogenesis in particular. However, the question of which miRNAs are specific to the chondrocyte phenotype has not been fully addressed. Using microarray analysis of miRNA expression during mesenchymal stem cell chondrogenic differentiation and detailed examination of the role of essential differentiation factors, such as SOX9, TGF-β, and the cell condensation phase, we characterize the repertoire of specific miRNAs involved in chondrocyte development, highlighting in particular miR-140 and miR-455. Further with the use of mRNA microarray data we integrate miRNA expression and mRNA expression during chondrogenesis to underline the particular importance of miR-140, especially the -5p strand. We provide a detailed identification and validation of direct targets of miR-140-5p in both chondrogenesis and adult chondrocytes with the use of microarray and 3'UTR analysis. This emphasizes the diverse array of targets and pathways regulated by miR-140-5p. We are also able to confirm previous experimentally identified targets but, additionally, identify a novel positive regulation of the Wnt signaling pathway by miR-140-5p. Wnt signaling has a complex role in chondrogenesis and skeletal development and these findings illustrate a previously unidentified role for miR-140-5p in regulation of Wnt signaling in these processes. Together these developments further highlight the role of miRNAs during chondrogenesis to improve our understanding of chondrocyte development and guide cartilage tissue engineering.
Keywords: Chondrogenesis; Differentiation; Epigenetics; Gene expression; Mesenchymal stem cells; miRNA.
© 2015 AlphaMed Press.
Figures

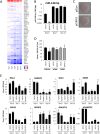

References
-
- Onyekwelu I, Goldring MB, Hidaka C. Chondrogenesis, joint formation, and articular cartilage regeneration. J Cell Biochem 2009;107:383–392. - PubMed
-
- Akiyama H. Control of chondrogenesis by the transcription factor Sox9. Mod Rheumatol 2008;18:213–219. - PubMed
-
- Murdoch AD, Grady LM, Ablett MP et al. Chondrogenic differentiation of human bone marrow stem cells in transwell cultures: Generation of scaffold‐free cartilage. Stem Cells 2007;25:2786–2796. - PubMed
-
- Pittenger MF, Mackay AM, Beck SC et al. Multilineage potential of adult human mesenchymal stem cells. Science 1999;284:143–147. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

