Effect of point mutations on Herbaspirillum seropedicae NifA activity
- PMID: 26176311
- PMCID: PMC4541686
- DOI: 10.1590/1414-431X20154522
Effect of point mutations on Herbaspirillum seropedicae NifA activity
Abstract
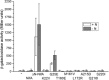
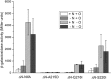
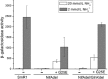
NifA is the transcriptional activator of the nif genes in Proteobacteria. It is usually regulated by nitrogen and oxygen, allowing biological nitrogen fixation to occur under appropriate conditions. NifA proteins have a typical three-domain structure, including a regulatory N-terminal GAF domain, which is involved in control by fixed nitrogen and not strictly required for activity, a catalytic AAA+ central domain, which catalyzes open complex formation, and a C-terminal domain involved in DNA-binding. In Herbaspirillum seropedicae, a β-proteobacterium capable of colonizing Graminae of agricultural importance, NifA regulation by ammonium involves its N-terminal GAF domain and the signal transduction protein GlnK. When the GAF domain is removed, the protein can still activate nif genes transcription; however, ammonium regulation is lost. In this work, we generated eight constructs resulting in point mutations in H. seropedicae NifA and analyzed their effect on nifH transcription in Escherichia coli and H. seropedicae. Mutations K22V, T160E, M161V, L172R, and A215D resulted in inactive proteins. Mutations Q216I and S220I produced partially active proteins with activity control similar to wild-type NifA. However, mutation G25E, located in the GAF domain, resulted in an active protein that did not require GlnK for activity and was partially sensitive to ammonium. This suggested that G25E may affect the negative interaction between the N-terminal GAF domain and the catalytic central domain under high ammonium concentrations, thus rendering the protein constitutively active, or that G25E could lead to a conformational change comparable with that when GlnK interacts with the GAF domain.
Figures

Similar articles
-
Interaction of GlnK with the GAF domain of Herbaspirillum seropedicae NifA mediates NH₄⁺-regulation.Biochimie. 2012 Apr;94(4):1041-7. doi: 10.1016/j.biochi.2012.01.007. Epub 2012 Jan 15. Biochimie. 2012. PMID: 22269934
-
Role of conserved cysteine residues in Herbaspirillum seropedicae NifA activity.Res Microbiol. 2009 Jul-Aug;160(6):389-95. doi: 10.1016/j.resmic.2009.06.002. Epub 2009 Jun 30. Res Microbiol. 2009. PMID: 19573596
-
In-trans regulation of the N-truncated-NIFA protein of Herbaspirillum seropedicae by the N-terminal domain.FEMS Microbiol Lett. 1999 Nov 15;180(2):157-61. doi: 10.1111/j.1574-6968.1999.tb08790.x. FEMS Microbiol Lett. 1999. PMID: 10556706
-
Regulation of nitrogen fixation in Klebsiella pneumoniae and Azotobacter vinelandii: NifL, transducing two environmental signals to the nif transcriptional activator NifA.J Mol Microbiol Biotechnol. 2002 May;4(3):235-42. J Mol Microbiol Biotechnol. 2002. PMID: 11931553 Review.
-
Azospirillum, a free-living nitrogen-fixing bacterium closely associated with grasses: genetic, biochemical and ecological aspects.FEMS Microbiol Rev. 2000 Oct;24(4):487-506. doi: 10.1111/j.1574-6976.2000.tb00552.x. FEMS Microbiol Rev. 2000. PMID: 10978548 Review.
Cited by
-
Regulation of Nitrogen Fixation in Bradyrhizobium sp. Strain DOA9 Involves Two Distinct NifA Regulatory Proteins That Are Functionally Redundant During Symbiosis but Not During Free-Living Growth.Front Microbiol. 2018 Jul 24;9:1644. doi: 10.3389/fmicb.2018.01644. eCollection 2018. Front Microbiol. 2018. PMID: 30087663 Free PMC article.
-
Engineering Posttranslational Regulation of Glutamine Synthetase for Controllable Ammonia Production in the Plant Symbiont Azospirillum brasilense.Appl Environ Microbiol. 2021 Jun 25;87(14):e0058221. doi: 10.1128/AEM.00582-21. Epub 2021 Jun 25. Appl Environ Microbiol. 2021. PMID: 33962983 Free PMC article.
References
-
- Huergo LF, Chandra G, Merrick M. P(II) signal transduction proteins: nitrogen regulation and beyond. FEMS Microbiol Rev. 2013;37:251–283. - PubMed
-
- Baldani JI, Baldani VLD, Seldin L, Dobereiner J. Characterization of Herbaspirillum seropedicae gen. nov., sp. nov., a root-associated nitrogen-fixing bacterium. Int J Syst Bacteriol. 1986;36:86–93. doi: 10.1099/00207713-36-1-86. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

