Tuning of Pectin Methylesterification: PECTIN METHYLESTERASE INHIBITOR 7 MODULATES THE PROCESSIVE ACTIVITY OF CO-EXPRESSED PECTIN METHYLESTERASE 3 IN A pH-DEPENDENT MANNER
- PMID: 26183897
- PMCID: PMC4645611
- DOI: 10.1074/jbc.M115.639534
Tuning of Pectin Methylesterification: PECTIN METHYLESTERASE INHIBITOR 7 MODULATES THE PROCESSIVE ACTIVITY OF CO-EXPRESSED PECTIN METHYLESTERASE 3 IN A pH-DEPENDENT MANNER
Abstract
Pectin methylesterases (PMEs) catalyze the demethylesterification of homogalacturonan domains of pectin in plant cell walls and are regulated by endogenous pectin methylesterase inhibitors (PMEIs). In Arabidopsis dark-grown hypocotyls, one PME (AtPME3) and one PMEI (AtPMEI7) were identified as potential interacting proteins. Using RT-quantitative PCR analysis and gene promoter::GUS fusions, we first showed that AtPME3 and AtPMEI7 genes had overlapping patterns of expression in etiolated hypocotyls. The two proteins were identified in hypocotyl cell wall extracts by proteomics. To investigate the potential interaction between AtPME3 and AtPMEI7, both proteins were expressed in a heterologous system and purified by affinity chromatography. The activity of recombinant AtPME3 was characterized on homogalacturonans (HGs) with distinct degrees/patterns of methylesterification. AtPME3 showed the highest activity at pH 7.5 on HG substrates with a degree of methylesterification between 60 and 80% and a random distribution of methyl esters. On the best HG substrate, AtPME3 generates long non-methylesterified stretches and leaves short highly methylesterified zones, indicating that it acts as a processive enzyme. The recombinant AtPMEI7 and AtPME3 interaction reduces the level of demethylesterification of the HG substrate but does not inhibit the processivity of the enzyme. These data suggest that the AtPME3·AtPMEI7 complex is not covalently linked and could, depending on the pH, be alternately formed and dissociated. Docking analysis indicated that the inhibition of AtPME3 could occur via the interaction of AtPMEI7 with a PME ligand-binding cleft structure. All of these data indicate that AtPME3 and AtPMEI7 could be partners involved in the fine tuning of HG methylesterification during plant development.
Keywords: degree of blockiness; gel diffusion assay; homology modeling; microscale thermophoresis; pectin methylesterase (PME); pectin methylesterase inhibitor (PMEI); plant biochemistry; plant cell wall; protein expression; protein-protein interaction.
© 2015 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures
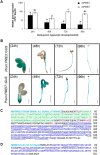
 ), and 3:1 (▨). Results are means ± S.D. of six replicates. The different letters indicate data sets significantly different according to Tukey's range test, preceded by a one-way ANOVA having p < 0.001.
), and 3:1 (▨). Results are means ± S.D. of six replicates. The different letters indicate data sets significantly different according to Tukey's range test, preceded by a one-way ANOVA having p < 0.001.
 ), 1:1 (
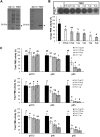
), 1:1 ( ), 1:2 (heavily dotted bars), 1:4 (□), and 1:8 (gray bars). Results are means ± S.D. (error bars) of six replicates. The different letters indicate data sets significantly different according to Tukey's range test, preceded by a one-way ANOVA having p < 0.001. At the most acidic pH, the maximum inhibition of PME3-His6 activity is reached for a ratio of ±60 pmol of PME3-His6/60 pmol of PMEI7-His6. B, inhibition of PME3-His6 activity by PMEI7-His6 on the HG96B82 substrate. PME3-His6 and PMEI7-His6 were preincubated for 30 min at 30 °C at pH 6.0. HG96B82 was added to the mixture and incubated for 30 min at 30 °C. The reaction was stopped at 90 °C for 10 min. PME activity was determined using a procedure adapted from Ref. . 1 μg of PME3-His6 was used with either PMEI7-His6 conservation solution (■) or a PME3-His6 activity/PMEI7-His6 (μg) ratio of 1:0.5 (
), 1:2 (heavily dotted bars), 1:4 (□), and 1:8 (gray bars). Results are means ± S.D. (error bars) of six replicates. The different letters indicate data sets significantly different according to Tukey's range test, preceded by a one-way ANOVA having p < 0.001. At the most acidic pH, the maximum inhibition of PME3-His6 activity is reached for a ratio of ±60 pmol of PME3-His6/60 pmol of PMEI7-His6. B, inhibition of PME3-His6 activity by PMEI7-His6 on the HG96B82 substrate. PME3-His6 and PMEI7-His6 were preincubated for 30 min at 30 °C at pH 6.0. HG96B82 was added to the mixture and incubated for 30 min at 30 °C. The reaction was stopped at 90 °C for 10 min. PME activity was determined using a procedure adapted from Ref. . 1 μg of PME3-His6 was used with either PMEI7-His6 conservation solution (■) or a PME3-His6 activity/PMEI7-His6 (μg) ratio of 1:0.5 ( ), 1:1 (▨), 1:2 (heavily dotted box), or 1:4 (□). Results are the means ± S.D. of two replicates.
), 1:1 (▨), 1:2 (heavily dotted box), or 1:4 (□). Results are the means ± S.D. of two replicates.
References
-
- Carpita N. C., Gibeaut D. M. (1993) Structural models of primary cell walls in flowering plants: consistency of molecular structure with the physical properties of the walls during growth. Plant J. 3, 1–30 - PubMed
-
- Caffall K. H., Mohnen D. (2009) The structure, function, and biosynthesis of plant cell wall pectic polysaccharides. Carbohydr. Res. 344, 1879–1900 - PubMed
-
- Ralet M.-C., Cabrera J. C., Bonnin E., Quéméner B., Hellìn P., Thibault J.-F. (2005) Mapping sugar beet pectin acetylation pattern. Phytochemistry 66, 1832–1843 - PubMed
-
- Pelloux J., Rustérucci C., Mellerowicz E. J. (2007) New insights into pectin methylesterase structure and function. Trends Plant Sci. 12, 267–277 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

