The Role of Metabolomics in Brain Metabolism Research
- PMID: 26201839
- PMCID: PMC4546538
- DOI: 10.1007/s11481-015-9621-1
The Role of Metabolomics in Brain Metabolism Research
Abstract
This special edition of the Journal of Neuroimmune Pharmacology focuses on the leading edge of metabolomics in brain metabolism research. The topics covered include a metabolomic field overview and the challenges in neuroscience metabolomics. The workflow and utility of different analytical platforms to profile complex biological matrices that include biofluids, brain tissue and cells, are shown in several case studies. These studies demonstrate how global and targeted metabolite profiling can be applied to distinguish disease stages and to understand the effects of drug action on the central nervous system (CNS). Finally, we discuss the importance of metabolomics to advance the understanding of brain function that includes ligand-receptor interactions and new insights into the mechanisms of CNS disorders.
Figures
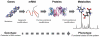



References
-
- Dickens Mountfort A, Larkin JR, Griffin JL, Davis BG, Claridge TDW, Sibson NR, Anthony DC. NMR-based metabolomics separates the distinct stages of disease in a chronic relapsing model of multiple sclerosis. J Neuroimmune Pharmacol. 2015 doi:10.1007/s11481-015-9622-0. - PubMed
-
- Duarte JM, Lei H, Mlynarik V, Gruetter R. The neurochemical profile quantified by in vivo 1H NMR spectroscopy. Neuroimage. 2012;61:342–362. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

