Characterization of the honeybee AmNaV1 channel and tools to assess the toxicity of insecticides
- PMID: 26202396
- PMCID: PMC4894402
- DOI: 10.1038/srep12475
Characterization of the honeybee AmNaV1 channel and tools to assess the toxicity of insecticides
Abstract
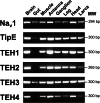
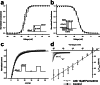
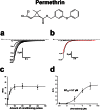
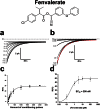
Pollination is important for both agriculture and biodiversity. For a significant number of plants, this process is highly, and sometimes exclusively, dependent on the pollination activity of honeybees. The large numbers of honeybee colony losses reported in recent years have been attributed to colony collapse disorder. Various hypotheses, including pesticide overuse, have been suggested to explain the disorder. Using the Xenopus oocytes expression system and two microelectrode voltage-clamp, we report the functional expression and the molecular, biophysical, and pharmacological characterization of the western honeybee's sodium channel (Apis Mellifera NaV1). The NaV1 channel is the primary target for pyrethroid insecticides in insect pests. We further report that the honeybee's channel is also sensitive to permethrin and fenvalerate, respectively type I and type II pyrethroid insecticides. Molecular docking of these insecticides revealed a binding site that is similar to sites previously identified in other insects. We describe in vitro and in silico tools that can be used to test chemical compounds. Our findings could be used to assess the risks that current and next generation pesticides pose to honeybee populations.
Figures

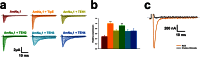


Similar articles
-
Molecular basis of selective resistance of the bumblebee BiNav1 sodium channel to tau-fluvalinate.Proc Natl Acad Sci U S A. 2017 Dec 5;114(49):12922-12927. doi: 10.1073/pnas.1711699114. Epub 2017 Nov 20. Proc Natl Acad Sci U S A. 2017. PMID: 29158414 Free PMC article.
-
Biophysical characterization of the Varroa destructor NaV1 sodium channel and its affinity for τ-fluvalinate insecticide.FASEB J. 2017 Jul;31(7):3066-3071. doi: 10.1096/fj.201601338R. Epub 2017 Mar 29. FASEB J. 2017. PMID: 28356346
-
A use-dependent sodium current modification induced by type I pyrethroid insecticides in honeybee antennal olfactory receptor neurons.Neurotoxicology. 2011 Jun;32(3):320-30. doi: 10.1016/j.neuro.2011.02.007. Epub 2011 Mar 4. Neurotoxicology. 2011. PMID: 21377491
-
Molecular Mechanism of Action and Selectivity of Sodium Ch annel Blocker Insecticides.Curr Med Chem. 2017;24(27):2912-2924. doi: 10.2174/0929867323666161216143844. Curr Med Chem. 2017. PMID: 27993108 Free PMC article. Review.
-
Neurotoxic actions of pyrethroid insecticides.Annu Rev Entomol. 1989;34:77-96. doi: 10.1146/annurev.en.34.010189.000453. Annu Rev Entomol. 1989. PMID: 2539040 Review.
Cited by
-
Honey bees long-lasting locomotor deficits after exposure to the diamide chlorantraniliprole are accompanied by brain and muscular calcium channels alterations.Sci Rep. 2019 Feb 15;9(1):2153. doi: 10.1038/s41598-019-39193-3. Sci Rep. 2019. PMID: 30770849 Free PMC article.
-
Molecular basis of selective resistance of the bumblebee BiNav1 sodium channel to tau-fluvalinate.Proc Natl Acad Sci U S A. 2017 Dec 5;114(49):12922-12927. doi: 10.1073/pnas.1711699114. Epub 2017 Nov 20. Proc Natl Acad Sci U S A. 2017. PMID: 29158414 Free PMC article.
-
A Locomotor Deficit Induced by Sublethal Doses of Pyrethroid and Neonicotinoid Insecticides in the Honeybee Apis mellifera.PLoS One. 2015 Dec 14;10(12):e0144879. doi: 10.1371/journal.pone.0144879. eCollection 2015. PLoS One. 2015. PMID: 26659095 Free PMC article.
-
Eukaryotic Voltage-Gated Sodium Channels: On Their Origins, Asymmetries, Losses, Diversification and Adaptations.Front Physiol. 2018 Nov 21;9:1406. doi: 10.3389/fphys.2018.01406. eCollection 2018. Front Physiol. 2018. PMID: 30519187 Free PMC article.
-
Cloning, functional expression, and pharmacological characterization of inwardly rectifying potassium channels (Kir) from Apis mellifera.Sci Rep. 2024 Apr 3;14(1):7834. doi: 10.1038/s41598-024-58234-0. Sci Rep. 2024. PMID: 38570597 Free PMC article.
References
-
- Morse R. A. & Calderone N. W. The value of honey bees as pollinators of U.S. crops in 2000. Bee Culture 128, 1–15 (2000).
-
- Gallai N., Salles J.-M., Settele J. & Vaissière B. E. Economic valuation of the vulnerability of world agriculture confronted with pollinator decline. Ecol. Econ. 68, 810–821 (2009).
-
- Carreck N. & Neumann P. Honey bee colony losses. J. Apicult. Res. 49, 1–1, 10.3896/IBRA.1.49.1.01 (2010). - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

