Structural prerequisites for G-protein activation by the neurotensin receptor
- PMID: 26205105
- PMCID: PMC4515772
- DOI: 10.1038/ncomms8895
Structural prerequisites for G-protein activation by the neurotensin receptor
Abstract
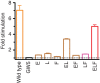
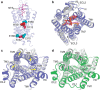
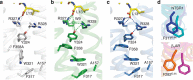
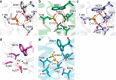
We previously determined the structure of neurotensin receptor NTSR1 in an active-like conformation with six thermostabilizing mutations bound to the peptide agonist neurotensin. This receptor was unable to activate G proteins, indicating that the mutations restricted NTSR1 to relate agonist binding to G-protein activation. Here we analyse the effect of three of those mutations (E166A(3.49), L310A(6.37), F358A(7.42)) and present two structures of NTSR1 able to catalyse nucleotide exchange at Gα. The presence of F358(7.42) causes the conserved W321(6.48) to adopt a side chain orientation parallel to the lipid bilayer sealing the collapsed Na(+) ion pocket and linking the agonist with residues in the lower receptor part implicated in GPCR activation. In the intracellular receptor half, the bulkier L310(6.37) side chain dictates the position of R167(3.50) of the highly conserved D/ERY motif. These residues, together with the presence of E166(3.49) provide determinants for G-protein activation by NTSR1.
Figures


Similar articles
-
Effect of thermostable mutations on the neurotensin receptor 1 (NTSR1) activation state.J Biomol Struct Dyn. 2020 Feb;38(2):340-353. doi: 10.1080/07391102.2019.1573705. Epub 2019 Feb 14. J Biomol Struct Dyn. 2020. PMID: 30706767
-
Structure and dynamics of a constitutively active neurotensin receptor.Sci Rep. 2016 Dec 7;6:38564. doi: 10.1038/srep38564. Sci Rep. 2016. PMID: 27924846 Free PMC article.
-
Thermostabilization of the neurotensin receptor NTS1.J Mol Biol. 2009 Jul 10;390(2):262-77. doi: 10.1016/j.jmb.2009.04.068. Epub 2009 May 5. J Mol Biol. 2009. PMID: 19422831 Free PMC article.
-
Maintaining cell sensitivity to G-protein coupled receptor agonists: neurotensin and the role of receptor gene activation.J Neuroendocrinol. 2001 May;13(5):473-9. doi: 10.1046/j.1365-2826.2001.00658.x. J Neuroendocrinol. 2001. PMID: 11328459 Review.
-
Phylogenetic history, pharmacological features, and signal transduction of neurotensin receptors in vertebrates.Ann N Y Acad Sci. 2009 Apr;1163:169-78. doi: 10.1111/j.1749-6632.2008.03636.x. Ann N Y Acad Sci. 2009. PMID: 19456337 Review.
Cited by
-
Probing the Conformation States of Neurotensin Receptor 1 Variants by NMR Site-Directed Methyl Labeling.Chembiochem. 2021 Jan 5;22(1):139-146. doi: 10.1002/cbic.202000541. Epub 2020 Nov 6. Chembiochem. 2021. PMID: 32881260 Free PMC article.
-
Structure of the neurotensin receptor 1 in complex with β-arrestin 1.Nature. 2020 Mar;579(7798):303-308. doi: 10.1038/s41586-020-1953-1. Epub 2020 Jan 16. Nature. 2020. PMID: 31945771 Free PMC article.
-
G protein-coupled receptors: the evolution of structural insight.AIMS Biophys. 2017;4(3):491-527. doi: 10.3934/biophy.2017.3.491. Epub 2017 Aug 21. AIMS Biophys. 2017. PMID: 29951585 Free PMC article.
-
Insights on the G protein-coupled receptor helix 8 solution structure and orientation using a neurotensin receptor 1 peptide.Protein Sci. 2024 Jun;33(6):e4976. doi: 10.1002/pro.4976. Protein Sci. 2024. PMID: 38757374 Free PMC article.
-
Screening of Membrane Protein Production by Comparison of Transient Expression in Insect and Mammalian Cells.Biomolecules. 2023 May 11;13(5):817. doi: 10.3390/biom13050817. Biomolecules. 2023. PMID: 37238687 Free PMC article.
References
-
- Choe H. W. et al. Crystal structure of metarhodopsin II. Nature 471, 651–655 (2011). - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

