Characterization of Dye-decolorizing Peroxidase (DyP) from Thermomonospora curvata Reveals Unique Catalytic Properties of A-type DyPs
- PMID: 26205819
- PMCID: PMC4645587
- DOI: 10.1074/jbc.M115.658807
Characterization of Dye-decolorizing Peroxidase (DyP) from Thermomonospora curvata Reveals Unique Catalytic Properties of A-type DyPs
Abstract
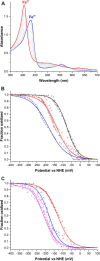
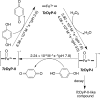
Dye-decolorizing peroxidases (DyPs) comprise a new family of heme peroxidases, which has received much attention due to their potential applications in lignin degradation. A new DyP from Thermomonospora curvata (TcDyP) was identified and characterized. Unlike other A-type enzymes, TcDyP is highly active toward a wide range of substrates including model lignin compounds, in which the catalytic efficiency with ABTS (kcat(app)/Km(app) = (1.7 × 10(7)) m(-1) s(-1)) is close to that of fungal DyPs. Stopped-flow spectroscopy was employed to elucidate the transient intermediates as well as the catalytic cycle involving wild-type (wt) and mutant TcDyPs. Although residues Asp(220) and Arg(327) are found necessary for compound I formation, His(312) is proposed to play roles in compound II reduction. Transient kinetics of hydroquinone (HQ) oxidation by wt-TcDyP showed that conversion of the compound II to resting state is a rate-limiting step, which will explain the contradictory observation made with the aspartate mutants of A-type DyPs. Moreover, replacement of His(312) and Arg(327) has significant effects on the oligomerization and redox potential (E°') of the enzyme. Both mutants were found to promote the formation of dimeric state and to shift E°' to a more negative potential. Not only do these results reveal the unique catalytic property of the A-type DyPs, but they will also facilitate the development of these enzymes as lignin degraders.
Keywords: dye-decolorizing peroxidase; enzyme kinetics; heme; lignin degradation; oligomerization; oxidation-reduction (redox); stopped-flow spectroscopy.
© 2015 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures







References
-
- Paliwal R., Rawat A. P., Rawat M., Rai J. P. (2012) Bioligninolysis: recent updates for biotechnological solution. Appl. Biochem. Biotechnol. 167, 1865–1889 - PubMed
-
- FitzPatrick M., Champagne P., Cunningham M. F., Whitney R. A. (2010) A biorefinery processing perspective: treatment of lignocellulosic materials for the production of value-added products. Bioresour. Technol. 101, 8915–8922 - PubMed
-
- Banerjee S., Mudliar S., Sen R., Giri B., Satpute D., Chakrabarti T., Pandey R. A. (2010) Commercializing lignocellulosic bioethanol: technology bottlenecks and possible remedies. Biofuels Bioprod. Bioref. 4, 77–93
-
- ten Have R., Teunissen P. J. (2001) Oxidative mechanisms involved in lignin degradation by white-rot fungi. Chem. Rev. 101, 3397–3413 - PubMed
-
- Brown M. E., Chang M. C. (2014) Exploring bacterial lignin degradation. Curr. Opin. Chem. Biol. 19, 1–7 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

