The Xis2d protein of CTnDOT binds to the intergenic region between the mob and tra operons
- PMID: 26212728
- PMCID: PMC4600444
- DOI: 10.1016/j.plasmid.2015.07.002
The Xis2d protein of CTnDOT binds to the intergenic region between the mob and tra operons
Abstract
CTnDOT is a 65kbp integrative and conjugative element (ICE) that carries genes encoding both tetracycline and erythromycin resistances. The excision operon of this element encodes Xis2c, Xis2d, and Exc proteins involved in the excision of CTnDOT from host chromosomes. These proteins are also required in the complex transcriptional regulation of the divergently transcribed transfer (tra) and mobilization (mob) operons of CTnDOT. Transcription of the tra operon is positively regulated by Xis2c and Xis2d, whereas, transcription of the mob operon is positively regulated by Xis2d and Exc. Xis2d is the only protein that is involved in the excision reaction, as well as the transcriptional regulation of both the mob and tra operons. This paper helps establish how Xis2d binds the DNA in the mob and tra region. Unlike other excisionase proteins, Xis2d binds a region of dyad symmetry. The binding site is located in the intergenic region between the mob and tra promoters, and once bound Xis2d induces a bend in the DNA. Xis2d binding to this region could be the preliminary step for the activation of both operons. Then the other proteins, like Exc, can interact with Xis2d and form higher order complexes.
Keywords: Conjugative transposon; Integrative and conjugative element; Recombination directionality factor.
Copyright © 2015 Elsevier B.V. All rights reserved.
Figures
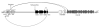







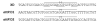

Similar articles
-
The excision proteins of CTnDOT positively regulate the transfer operon.Plasmid. 2013 Mar;69(2):172-9. doi: 10.1016/j.plasmid.2012.12.001. Epub 2012 Dec 10. Plasmid. 2013. PMID: 23237854 Free PMC article.
-
Tetracycline-related transcriptional regulation of the CTnDOT mobilization region.J Bacteriol. 2013 Dec;195(24):5431-8. doi: 10.1128/JB.00691-13. Epub 2013 Sep 27. J Bacteriol. 2013. PMID: 24078614 Free PMC article.
-
Interactions of the excision proteins of CTnDOT in the attR intasome.Plasmid. 2013 Sep;70(2):190-200. doi: 10.1016/j.plasmid.2013.03.009. Epub 2013 Apr 17. Plasmid. 2013. PMID: 23603449 Free PMC article.
-
The Integration and Excision of CTnDOT.Microbiol Spectr. 2015 Apr;3(2):MDNA3-0020-2014. doi: 10.1128/microbiolspec.MDNA3-0020-2014. Microbiol Spectr. 2015. PMID: 26104696 Free PMC article. Review.
-
Regulation of CTnDOT conjugative transfer is a complex and highly coordinated series of events.mBio. 2013 Oct 29;4(6):e00569-13. doi: 10.1128/mBio.00569-13. mBio. 2013. PMID: 24169574 Free PMC article. Review.
Cited by
-
Timing of integration into the chromosome is critical for the fitness of an integrative and conjugative element and its bacterial host.PLoS Genet. 2023 Feb 13;19(2):e1010524. doi: 10.1371/journal.pgen.1010524. eCollection 2023 Feb. PLoS Genet. 2023. PMID: 36780569 Free PMC article.
-
Elucidating the role of mycobacteriophage D29-encoded Gp36 in DNA binding and phage gene expression regulation.Nucleic Acids Res. 2025 Jul 8;53(13):gkaf662. doi: 10.1093/nar/gkaf662. Nucleic Acids Res. 2025. PMID: 40671521 Free PMC article.
-
The hidden life of integrative and conjugative elements.FEMS Microbiol Rev. 2017 Jul 1;41(4):512-537. doi: 10.1093/femsre/fux008. FEMS Microbiol Rev. 2017. PMID: 28369623 Free PMC article. Review.
References
-
- Salyers AA, Gupta A, Wang Y. Human intestinal bacteria as reservoirs for antibiotic resistance genes. Trends in microbiology. 2004;12:412–416. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

