One of Three Pex11 Family Members Is Required for Peroxisomal Proliferation and Full Virulence of the Rice Blast Fungus Magnaporthe oryzae
- PMID: 26218097
- PMCID: PMC4517885
- DOI: 10.1371/journal.pone.0134249
One of Three Pex11 Family Members Is Required for Peroxisomal Proliferation and Full Virulence of the Rice Blast Fungus Magnaporthe oryzae
Abstract
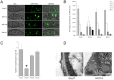
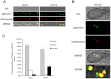
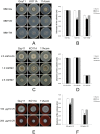
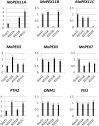
Peroxisomes play important roles in metabolisms of eukaryotes and infection of plant fungal pathogens. These organelles proliferate by de novo formation or division in response to environmental stimulation. Although the assembly of peroxisomes was documented in fungal pathogens, their division and its relationship to pathogenicity remain obscure. In present work, we analyzed the roles of three Pex11 family members in peroxisomal division and pathogenicity of the rice blast fungus Magnaporthe oryzae. Deletion of MoPEX11A led to fewer but enlarged peroxisomes, and impaired the separation of Woronin bodies from peroxisomes, while deletion of MoPEX11B or MoPEX11C put no evident impacts to peroxisomal profiles. MoPEX11A mutant exhibited typical peroxisome related defects, delayed conidial germination and appressoria formation, and decreased appressorial turgor and host penetration. As a result, the virulence of MoPEX11A mutant was greatly reduced. Deletion of MoPEX11B and MoPEX11C did not alter the virulence of the fungus. Further, double or triple deletions of the three genes were unable to enhance the virulence decrease in MoPEX11A mutant. Our data indicated that MoPEX11A is the main factor modulating peroxisomal division and is required for full virulence of the fungus.
Conflict of interest statement
Figures




Similar articles
-
Roles of Peroxisomes in the Rice Blast Fungus.Biomed Res Int. 2016;2016:9343417. doi: 10.1155/2016/9343417. Epub 2016 Aug 16. Biomed Res Int. 2016. PMID: 27610388 Free PMC article. Review.
-
Pex13 and Pex14, the key components of the peroxisomal docking complex, are required for peroxisome formation, host infection and pathogenicity-related morphogenesis in Magnaporthe oryzae.Virulence. 2019 Dec;10(1):292-314. doi: 10.1080/21505594.2019.1598172. Virulence. 2019. PMID: 30905264 Free PMC article.
-
Pex14/17, a filamentous fungus-specific peroxin, is required for the import of peroxisomal matrix proteins and full virulence of Magnaporthe oryzae.Mol Plant Pathol. 2017 Dec;18(9):1238-1252. doi: 10.1111/mpp.12487. Epub 2016 Dec 19. Mol Plant Pathol. 2017. PMID: 27571711 Free PMC article.
-
PTS1 peroxisomal import pathway plays shared and distinct roles to PTS2 pathway in development and pathogenicity of Magnaporthe oryzae.PLoS One. 2013;8(2):e55554. doi: 10.1371/journal.pone.0055554. Epub 2013 Feb 6. PLoS One. 2013. PMID: 23405169 Free PMC article.
-
The role of glycerol in the pathogenic lifestyle of the rice blast fungus Magnaporthe oryzae.Environ Microbiol. 2017 Mar;19(3):1008-1016. doi: 10.1111/1462-2920.13688. Epub 2017 Mar 1. Environ Microbiol. 2017. PMID: 28165657 Review.
Cited by
-
The peroxins BcPex8, BcPex10, and BcPex12 are required for the development and pathogenicity of Botrytis cinerea.Front Microbiol. 2022 Sep 6;13:962500. doi: 10.3389/fmicb.2022.962500. eCollection 2022. Front Microbiol. 2022. PMID: 36147853 Free PMC article.
-
Peroxin MoPex22 Regulates the Import of Peroxisomal Matrix Proteins and Appressorium-Mediated Plant Infection in Magnaporthe oryzae.J Fungi (Basel). 2024 Feb 10;10(2):143. doi: 10.3390/jof10020143. J Fungi (Basel). 2024. PMID: 38392815 Free PMC article.
-
Dynamic Regulation of Peroxisomes and Mitochondria during Fungal Development.J Fungi (Basel). 2020 Nov 20;6(4):302. doi: 10.3390/jof6040302. J Fungi (Basel). 2020. PMID: 33233491 Free PMC article. Review.
-
Roles of Peroxisomes in the Rice Blast Fungus.Biomed Res Int. 2016;2016:9343417. doi: 10.1155/2016/9343417. Epub 2016 Aug 16. Biomed Res Int. 2016. PMID: 27610388 Free PMC article. Review.
-
MoLrp1-mediated signaling induces nuclear accumulation of MoMsn2 to facilitate fatty acid oxidation for infectious growth of the rice blast fungus.Plant Commun. 2023 Jul 10;4(4):100561. doi: 10.1016/j.xplc.2023.100561. Epub 2023 Feb 11. Plant Commun. 2023. PMID: 36774535 Free PMC article.
References
-
- Vanacova S, Liston DR, Tachezy J, Johnson PJ (2003) Molecular biology of the amitochondriate parasites, Giardia intestinalis, Entamoeba histolytica and Trichomonas vaginalis . International Journal for Parasitology 33: 235–255. - PubMed
-
- Lazarow PB, Fujiki Y (1985) Biogenesis of peroxisomes. Annual Review of Cell Biology 1: 489–530. - PubMed
-
- Elgersma Y, Tabak HF (1996) Proteins involved in peroxisome biogenesis and functioning. Biochimica Et Biophysica Acta-Reviews on Biomembranes 1286: 269–283. - PubMed
-
- Vandenbosch H, Schrakamp G, Hardeman D, Zomer AWM, Wanders RJA, Schutgens RBH (1993) Ether lipid-synthesis and its deficiency in peroxisomal disorders. Biochimie 75: 183–189. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

