Development and applications of single-cycle infectious influenza A virus (sciIAV)
- PMID: 26220478
- PMCID: PMC4728073
- DOI: 10.1016/j.virusres.2015.07.013
Development and applications of single-cycle infectious influenza A virus (sciIAV)
Abstract
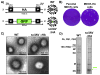
The diverse host range, high transmissibility, and rapid evolution of influenza A viruses justify the importance of containing pathogenic viruses studied in the laboratory. Other than physically or mechanically changing influenza A virus containment procedures, modifying the virus to only replicate for a single round of infection similarly ensures safety and consequently decreases the level of biosafety containment required to study highly pathogenic members in the virus family. This biological containment is more ideal because it is less apt to computer, machine, or human error. With many necessary proteins that can be deleted, generation of single-cycle infectious influenza A viruses (sciIAV) can be achieved using a variety of approaches. Here, we review the recent burst in sciIAV generation and summarize the applications and findings on this important human pathogen using biocontained viral mimics.
Keywords: Influenza HA-expressing MDCK cells (MDCK-HA); Influenza vaccine; Reporter virus; Reverse genetics; Single-cycle virus; Viral vectors.
Copyright © 2015 Elsevier B.V. All rights reserved.
Figures


Similar articles
-
Generation and Characterization of Single-Cycle Infectious Canine Influenza A Virus (sciCIV) and Its Use as Vaccine Platform.Methods Mol Biol. 2022;2465:227-255. doi: 10.1007/978-1-0716-2168-4_13. Methods Mol Biol. 2022. PMID: 35118625
-
Application of a Biologically Contained Reporter System To Study Gain-of-Function H5N1 Influenza A Viruses with Pandemic Potential.mSphere. 2020 Aug 26;5(4):e00423-20. doi: 10.1128/mSphere.00423-20. mSphere. 2020. PMID: 32848003 Free PMC article.
-
Improvement of influenza A/Fujian/411/02 (H3N2) virus growth in embryonated chicken eggs by balancing the hemagglutinin and neuraminidase activities, using reverse genetics.J Virol. 2005 Jun;79(11):6763-71. doi: 10.1128/JVI.79.11.6763-6771.2005. J Virol. 2005. PMID: 15890915 Free PMC article.
-
Enhancement of influenza virus transmission by gene reassortment.Curr Top Microbiol Immunol. 2014;385:185-204. doi: 10.1007/82_2014_389. Curr Top Microbiol Immunol. 2014. PMID: 25048543 Review.
-
Transmission of influenza A viruses.Virology. 2015 May;479-480:234-46. doi: 10.1016/j.virol.2015.03.009. Epub 2015 Mar 24. Virology. 2015. PMID: 25812763 Free PMC article. Review.
Cited by
-
Characterization of Influenza Virus Pseudotyped with Ebolavirus Glycoprotein.J Virol. 2018 Jan 30;92(4):e00941-17. doi: 10.1128/JVI.00941-17. Print 2018 Feb 15. J Virol. 2018. PMID: 29212933 Free PMC article.
-
Generation and Characterization of Single-Cycle Infectious Canine Influenza A Virus (sciCIV) and Its Use as Vaccine Platform.Methods Mol Biol. 2022;2465:227-255. doi: 10.1007/978-1-0716-2168-4_13. Methods Mol Biol. 2022. PMID: 35118625
-
Development of in vitro and in vivo neutralization assays based on the pseudotyped H7N9 virus.Sci Rep. 2018 May 31;8(1):8484. doi: 10.1038/s41598-018-26822-6. Sci Rep. 2018. PMID: 29855533 Free PMC article.
-
Advances in Development and Application of Influenza Vaccines.Front Immunol. 2021 Jul 13;12:711997. doi: 10.3389/fimmu.2021.711997. eCollection 2021. Front Immunol. 2021. PMID: 34326849 Free PMC article. Review.
-
Replication-competent fluorescent-expressing influenza B virus.Virus Res. 2016 Feb 2;213:69-81. doi: 10.1016/j.virusres.2015.11.014. Epub 2015 Nov 15. Virus Res. 2016. PMID: 26590325 Free PMC article.
References
-
- Wkly Epidemiol Rec; Assays for neutralizing antibody to influenza viruses. Report of an informal scientific workshop; Dresden. 18–19 March 2003; pp. 290–293. - PubMed
-
- Abe T, Fujimori T. Reporter mouse lines for fluorescence imaging. Dev Growth Differ. 2013;55(4):390–405. - PubMed
-
- Aoki H, Nishiyama Y, Tsurumi T, Shibata M, Ito Y, Seo H, Yoshii S, Maeno K. Mechanism of interference between influenza A/WSN and B/Kanagawa viruses. J Gen Virol. 1984;65(Pt 8):1385–1393. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

