Towards a comprehensive picture of alloacceptor tRNA remolding in metazoan mitochondrial genomes
- PMID: 26227972
- PMCID: PMC4783518
- DOI: 10.1093/nar/gkv746
Towards a comprehensive picture of alloacceptor tRNA remolding in metazoan mitochondrial genomes
Abstract
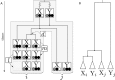
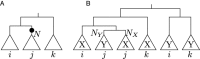
Remolding of tRNAs is a well-documented process in mitochondrial genomes that changes the identity of a tRNA. It involves a duplication of a tRNA gene, a mutation that changes the anticodon and the loss of the ancestral tRNA gene. The net effect is a functional tRNA that is more closely related to tRNAs of a different alloacceptor family than to tRNAs with the same anticodon in related species. Beyond being of interest for understanding mitochondrial tRNA function and evolution, tRNA remolding events can lead to artifacts in the annotation of mitogenomes and thus in studies of mitogenomic evolution. Therefore, it is important to identify and catalog these events. Here we describe novel methods to detect tRNA remolding in large-scale data sets and apply them to survey tRNA remolding throughout animal evolution. We identify several novel remolding events in addition to the ones previously mentioned in the literature. A detailed analysis of these remoldings showed that many of them are derived from ancestral events.
© The Author(s) 2015. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures




References
-
- McBride H.M., Neuspiel M., Wasiak S. Mitochondria: more than just a powerhouse. Curr. Biol. 2006;16:551–560. - PubMed
-
- Duchêne A.-M., Pujol C., Maréchal-Drouard L. Import of tRNAs and aminoacyl-tRNA synthetases into mitochondria. Curr. Genet. 2009;55:1–18. - PubMed
-
- Beagley C., Macfarlane J., Pont-Kingdon G., Okimoto R., Okada N., Wolstenholme D. Mitochondrial genomes of Anthozoa (Cnidaria) Prog. Cell Res. 1995;5:149–153.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

