Toolbox Approaches Using Molecular Markers and 16S rRNA Gene Amplicon Data Sets for Identification of Fecal Pollution in Surface Water
- PMID: 26231650
- PMCID: PMC4579440
- DOI: 10.1128/AEM.02032-15
Toolbox Approaches Using Molecular Markers and 16S rRNA Gene Amplicon Data Sets for Identification of Fecal Pollution in Surface Water
Abstract
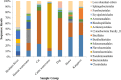
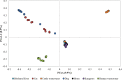
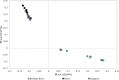
In this study, host-associated molecular markers and bacterial 16S rRNA gene community analysis using high-throughput sequencing were used to identify the sources of fecal pollution in environmental waters in Brisbane, Australia. A total of 92 fecal and composite wastewater samples were collected from different host groups (cat, cattle, dog, horse, human, and kangaroo), and 18 water samples were collected from six sites (BR1 to BR6) along the Brisbane River in Queensland, Australia. Bacterial communities in the fecal, wastewater, and river water samples were sequenced. Water samples were also tested for the presence of bird-associated (GFD), cattle-associated (CowM3), horse-associated, and human-associated (HF183) molecular markers, to provide multiple lines of evidence regarding the possible presence of fecal pollution associated with specific hosts. Among the 18 water samples tested, 83%, 33%, 17%, and 17% were real-time PCR positive for the GFD, HF183, CowM3, and horse markers, respectively. Among the potential sources of fecal pollution in water samples from the river, DNA sequencing tended to show relatively small contributions from wastewater treatment plants (up to 13% of sequence reads). Contributions from other animal sources were rarely detected and were very small (<3% of sequence reads). Source contributions determined via sequence analysis versus detection of molecular markers showed variable agreement. A lack of relationships among fecal indicator bacteria, host-associated molecular markers, and 16S rRNA gene community analysis data was also observed. Nonetheless, we show that bacterial community and host-associated molecular marker analyses can be combined to identify potential sources of fecal pollution in an urban river. This study is a proof of concept, and based on the results, we recommend using bacterial community analysis (where possible) along with PCR detection or quantification of host-associated molecular markers to provide information on the sources of fecal pollution in waterways.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures

Similar articles
-
Human-Associated Lachnospiraceae Genetic Markers Improve Detection of Fecal Pollution Sources in Urban Waters.Appl Environ Microbiol. 2018 Jul 2;84(14):e00309-18. doi: 10.1128/AEM.00309-18. Print 2018 Jul 15. Appl Environ Microbiol. 2018. PMID: 29728386 Free PMC article.
-
Quantitative identification of fecal water pollution sources by TaqMan real-time PCR assays using Bacteroidales 16S rRNA genetic markers.Appl Microbiol Biotechnol. 2010 Dec;88(6):1373-83. doi: 10.1007/s00253-010-2880-0. Epub 2010 Sep 25. Appl Microbiol Biotechnol. 2010. PMID: 20871990
-
Evaluation of host-specific Bacteroidales 16S rRNA gene markers as a complementary tool for detecting fecal pollution in a prairie watershed.Water Res. 2009 Nov;43(19):4838-49. doi: 10.1016/j.watres.2009.06.045. Epub 2009 Jun 27. Water Res. 2009. PMID: 19604534
-
Fecal pollution source tracking toolbox for identification, evaluation and characterization of fecal contamination in receiving urban surface waters and groundwater.Sci Total Environ. 2015 Dec 15;538:38-57. doi: 10.1016/j.scitotenv.2015.07.155. Epub 2015 Aug 22. Sci Total Environ. 2015. PMID: 26298247 Review.
-
Fecal pollution: new trends and challenges in microbial source tracking using next-generation sequencing.Environ Microbiol. 2018 Sep;20(9):3132-3140. doi: 10.1111/1462-2920.14281. Epub 2018 Aug 5. Environ Microbiol. 2018. PMID: 29797757 Review.
Cited by
-
A Community Multi-Omics Approach towards the Assessment of Surface Water Quality in an Urban River System.Int J Environ Res Public Health. 2017 Mar 14;14(3):303. doi: 10.3390/ijerph14030303. Int J Environ Res Public Health. 2017. PMID: 28335448 Free PMC article.
-
Quantifying the contribution of microbial immigration in engineered water systems.Microbiome. 2019 Nov 6;7(1):144. doi: 10.1186/s40168-019-0760-0. Microbiome. 2019. PMID: 31694700 Free PMC article. Review.
-
Coupling growth kinetics modeling with machine learning reveals microbial immigration impacts and identifies key environmental parameters in a biological wastewater treatment process.Microbiome. 2019 Apr 17;7(1):65. doi: 10.1186/s40168-019-0682-x. Microbiome. 2019. PMID: 30995941 Free PMC article.
-
The Use of Ribosomal RNA as a Microbial Source Tracking Target Highlights the Assay Host-Specificity Requirement in Water Quality Assessments.Front Microbiol. 2021 Jun 2;12:673306. doi: 10.3389/fmicb.2021.673306. eCollection 2021. Front Microbiol. 2021. PMID: 34149662 Free PMC article.
-
Microbial Indicators of Fecal Pollution: Recent Progress and Challenges in Assessing Water Quality.Curr Environ Health Rep. 2020 Sep;7(3):311-324. doi: 10.1007/s40572-020-00278-1. Curr Environ Health Rep. 2020. PMID: 32542574 Free PMC article. Review.
References
-
- Tallon P, Magajna B, Lofranco C, Leung KT. 2005. Microbial indicators of faecal contamination in water: a current perspective. Water Air Soil Pollut 166:139–166. doi:10.1007/s11270-005-7905-4. - DOI
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

