NagC represses N-acetyl-glucosamine utilization genes in Vibrio fischeri within the light organ of Euprymna scolopes
- PMID: 26236308
- PMCID: PMC4505101
- DOI: 10.3389/fmicb.2015.00741
NagC represses N-acetyl-glucosamine utilization genes in Vibrio fischeri within the light organ of Euprymna scolopes
Abstract
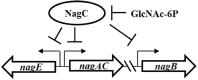
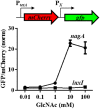
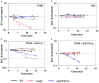
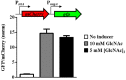
Bacteria often use transcription factors to regulate the expression of metabolic genes in accordance to available nutrients. NagC is a repressor conserved among γ-proteobacteria that regulates expression of enzymes involved in the metabolism of N-acetyl-glucosamine (GlcNAc). The polymeric form of GlcNAc, known as chitin, has been shown to play roles in chemotactic signaling and nutrition within the light organ symbiosis established between the marine bacterium Vibrio fischeri and the Hawaiian squid Euprymna scolopes. Here, we investigate the impact of NagC regulation on the physiology of V. fischeri. We find that NagC repression contributes to the fitness of V. fischeri in the absence of GlcNAc. In addition, the inability to de-repress expression of NagC-regulated genes reduces the fitness of V. fischeri in the presence of GlcNAc. We find that chemotaxis toward GlcNAc or chitobiose, a dimeric form of GlcNAc, is independent of NagC regulation. Finally, we show that NagC represses gene expression during the early stages of symbiosis. Our data suggest that the ability to regulate gene expression with NagC contributes to the overall fitness of V. fischeri in environments that vary in levels of GlcNAc. Furthermore, our finding that NagC represses gene expression within the squid light organ during an early stage of symbiosis supports the notion that the ability of the squid to provide a source of GlcNAc emerges later in host development.
Keywords: Euprymna scolopes; N-acetyl-glucosamine; NagC; Vibrio; symbiosis.
Figures

Similar articles
-
The N-acetyl-D-glucosamine repressor NagC of Vibrio fischeri facilitates colonization of Euprymna scolopes.Mol Microbiol. 2011 Nov;82(4):894-903. doi: 10.1111/j.1365-2958.2011.07858.x. Epub 2011 Oct 12. Mol Microbiol. 2011. PMID: 21992506 Free PMC article.
-
Squid-derived chitin oligosaccharides are a chemotactic signal during colonization by Vibrio fischeri.Appl Environ Microbiol. 2012 Jul;78(13):4620-6. doi: 10.1128/AEM.00377-12. Epub 2012 Apr 20. Appl Environ Microbiol. 2012. PMID: 22522684 Free PMC article.
-
Aposymbiotic culture of the sepiolid squid Euprymna scolopes: role of the symbiotic bacterium Vibrio fischeri in host animal growth, development, and light organ morphogenesis.J Exp Zool. 2000 Feb 15;286(3):280-96. J Exp Zool. 2000. PMID: 10653967
-
The Role of Hemocytes in the Hawaiian Bobtail Squid, Euprymna scolopes: A Model Organism for Studying Beneficial Host-Microbe Interactions.Front Microbiol. 2017 Jan 6;7:2013. doi: 10.3389/fmicb.2016.02013. eCollection 2016. Front Microbiol. 2017. PMID: 28111565 Free PMC article. Review.
-
Gimme shelter: how Vibrio fischeri successfully navigates an animal's multiple environments.Front Microbiol. 2013 Nov 29;4:356. doi: 10.3389/fmicb.2013.00356. eCollection 2013. Front Microbiol. 2013. PMID: 24348467 Free PMC article. Review.
Cited by
-
The NAG Sensor NagC Regulates LEE Gene Expression and Contributes to Gut Colonization by Escherichia coli O157:H7.Front Cell Infect Microbiol. 2017 Apr 24;7:134. doi: 10.3389/fcimb.2017.00134. eCollection 2017. Front Cell Infect Microbiol. 2017. PMID: 28484684 Free PMC article.
-
Rethinking the roles of CRP, cAMP, and sugar-mediated global regulation in the Vibrionaceae.Curr Genet. 2016 Feb;62(1):39-45. doi: 10.1007/s00294-015-0508-8. Epub 2015 Jul 28. Curr Genet. 2016. PMID: 26215147 Review.
-
Sulfur availability for Vibrio fischeri growth during symbiosis establishment depends on biogeography within the squid light organ.Mol Microbiol. 2019 Mar;111(3):621-636. doi: 10.1111/mmi.14177. Epub 2019 Jan 2. Mol Microbiol. 2019. PMID: 30506600 Free PMC article.
-
Intraspecific Competition Impacts Vibrio fischeri Strain Diversity during Initial Colonization of the Squid Light Organ.Appl Environ Microbiol. 2016 May 2;82(10):3082-91. doi: 10.1128/AEM.04143-15. Print 2016 May 15. Appl Environ Microbiol. 2016. PMID: 27016564 Free PMC article.
-
Niche-Specific Impact of a Symbiotic Function on the Persistence of Microbial Symbionts within a Natural Host.Appl Environ Microbiol. 2016 Sep 16;82(19):5990-6. doi: 10.1128/AEM.01770-16. Print 2016 Oct 1. Appl Environ Microbiol. 2016. PMID: 27474717 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

