Estimated herd prevalence and sequence types of Coxiella burnetii in bulk tank milk samples from commercial dairies in Indiana
- PMID: 26248712
- PMCID: PMC4528813
- DOI: 10.1186/s12917-015-0517-3
Estimated herd prevalence and sequence types of Coxiella burnetii in bulk tank milk samples from commercial dairies in Indiana
Abstract
Background: Coxiella burnetii is the etiologic agent of Q fever, a zoonotic disease causing influenza-like illness, pregnancy loss, cardiovascular disease and chronic fatigue syndrome in people. C. burnetii is considered to be enzootic in ruminants, but clinical signs of infection do not always manifest. National studies have documented the presence of C. burnetii in dairy herds in Indiana. This represents an opportunity to better characterize the distribution and prevalence of C. burnetii infection at the state scale, allowing evaluation of the need for surveillance and response planning to occur at this level. A cross-sectional study was conducted to estimate the herd prevalence of C. burnetii in commercial cattle dairies in Indiana and characterize the strains of C. burnetii within these dairies.
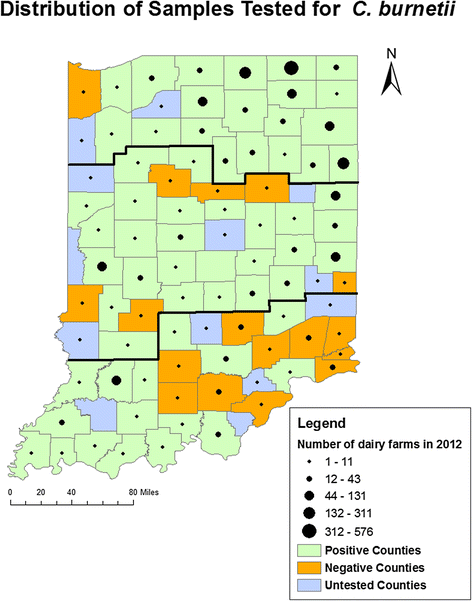
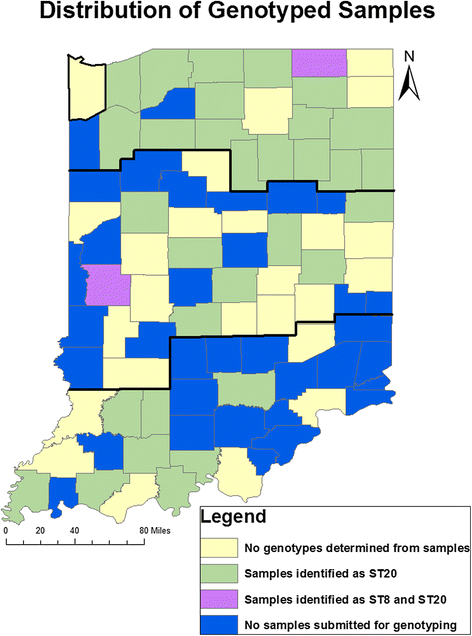
Results: Bulk tank milk samples were collected between June and August of 2011 by the Indiana State Board of Animal Health (ISBOAH). A total of 316 of these samples were tested for the IS1111 transposon of C. burnetii using quantitative real time polymerase chain reaction (PCR). Single nucleotide polymorphism (SNP) genotyping was used to identify the multispacer sequence genotypes (ST) present in samples where the IS1111 transposon was identified. The geographic distribution of dairies testing positive for C. burnetii DNA and the identified STs were also evaluated. The estimated overall herd prevalence for C. burnetii DNA was 61.1 % (95 % CI 55.6-66.3 %). The highest estimated regional prevalence was 70.2 % in the Central region of Indiana. An ST was identifiable in 74 of the positive 178 samples (41.6 %) and none of the 10 negative samples tested. Of these samples, 71 (95.9 %) were identified as ST20, 2 (2.7 %) as ST8 and a combination of ST20 and ST8 was identified in a single sample.
Conclusions: C. burnetii is present in dairy herds throughout Indiana. Indiana follows national trends with ST20 most commonly identified. The presence of multiple STs in a single bulk tank sample indicates that multiple strains of C. burnetii can circulate within a herd. This supports potential transmission of C. burnetii between goats and cattle, presenting the potential for a switch in the dominant genotype found in a given species.
Figures
References
-
- Derrick EH. Q fever, a new fever entity: clinical features, diagnosis and laboratory investigation. Med J Aust. 1937;2(8):281–99. - PubMed
-
- Hugh-Jones ME, Hubbert WT, Hagstad HV. Zoonoses: recognition, control and prevention. Ames Iowa: Iowa State University Press; 1995.
-
- McQuiston JH, Holman RC, McCall CL, Childs JE, Swerdlow DL, Thompson HA. National surveillance and the epidemiology of human Q fever in the United States, 1978–2004. Am J Trop Med Hyg. 2006;75(1):36–40. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

