Structure, Dynamics, and Allosteric Potential of Ionotropic Glutamate Receptor N-Terminal Domains
- PMID: 26255587
- PMCID: PMC4576161
- DOI: 10.1016/j.bpj.2015.06.061
Structure, Dynamics, and Allosteric Potential of Ionotropic Glutamate Receptor N-Terminal Domains
Abstract
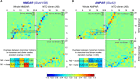
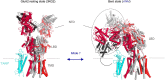
Ionotropic glutamate receptors (iGluRs) are tetrameric cation channels that mediate synaptic transmission and plasticity. They have a unique modular architecture with four domains: the intracellular C-terminal domain (CTD) that is involved in synaptic targeting, the transmembrane domain (TMD) that forms the ion channel, the membrane-proximal ligand-binding domain (LBD) that binds agonists such as L-glutamate, and the distal N-terminal domain (NTD), whose function is the least clear. The extracellular portion, comprised of the LBD and NTD, is loosely arranged, mediating complex allosteric regulation and providing a rich target for drug development. Here, we briefly review recent work on iGluR NTD structure and dynamics, and further explore the allosteric potential for the NTD in AMPA-type iGluRs using coarse-grained simulations. We also investigate mechanisms underlying the established NTD allostery in NMDA-type iGluRs, as well as the fold-related metabotropic glutamate and GABAB receptors. We show that the clamshell motions intrinsically favored by the NTD bilobate fold are coupled to dimeric and higher-order rearrangements that impact the iGluR LBD and ultimately the TMD. Finally, we explore the dynamics of intact iGluRs and describe how it might affect receptor operation in a synaptic environment.
Copyright © 2015 The Authors. Published by Elsevier Inc. All rights reserved.
Figures



Similar articles
-
Cooperative Dynamics of Intact AMPA and NMDA Glutamate Receptors: Similarities and Subfamily-Specific Differences.Structure. 2015 Sep 1;23(9):1692-1704. doi: 10.1016/j.str.2015.07.002. Epub 2015 Aug 6. Structure. 2015. PMID: 26256538 Free PMC article.
-
Comparative dynamics of NMDA- and AMPA-glutamate receptor N-terminal domains.Structure. 2012 Nov 7;20(11):1838-49. doi: 10.1016/j.str.2012.08.012. Epub 2012 Sep 6. Structure. 2012. PMID: 22959625 Free PMC article.
-
Allosteric coupling of sub-millisecond clamshell motions in ionotropic glutamate receptor ligand-binding domains.Commun Biol. 2021 Sep 9;4(1):1056. doi: 10.1038/s42003-021-02605-0. Commun Biol. 2021. PMID: 34504293 Free PMC article.
-
Activation and desensitization of ionotropic glutamate receptors by selectively triggering pre-existing motions.Neurosci Lett. 2019 May 1;700:22-29. doi: 10.1016/j.neulet.2018.02.050. Epub 2018 Feb 23. Neurosci Lett. 2019. PMID: 29481851 Free PMC article. Review.
-
Emerging structural insights into the function of ionotropic glutamate receptors.Trends Biochem Sci. 2015 Jun;40(6):328-37. doi: 10.1016/j.tibs.2015.04.002. Epub 2015 May 1. Trends Biochem Sci. 2015. PMID: 25941168 Free PMC article. Review.
Cited by
-
Design of Elastic Networks with Evolutionary Optimized Long-Range Communication as Mechanical Models of Allosteric Proteins.Biophys J. 2017 Aug 8;113(3):558-571. doi: 10.1016/j.bpj.2017.06.043. Biophys J. 2017. PMID: 28793211 Free PMC article.
-
Pharmacological characterisation of S 47445, a novel positive allosteric modulator of AMPA receptors.PLoS One. 2017 Sep 8;12(9):e0184429. doi: 10.1371/journal.pone.0184429. eCollection 2017. PLoS One. 2017. PMID: 28886144 Free PMC article.
-
InSty: A ProDy Module for Evaluating Protein Interactions and Stability.J Mol Biol. 2025 Aug 1;437(15):169009. doi: 10.1016/j.jmb.2025.169009. Epub 2025 Feb 13. J Mol Biol. 2025. PMID: 39954779
-
The Concise Guide to PHARMACOLOGY 2023/24: Ion channels.Br J Pharmacol. 2023 Oct;180 Suppl 2(Suppl 2):S145-S222. doi: 10.1111/bph.16178. Br J Pharmacol. 2023. PMID: 38123150 Free PMC article.
-
Dependence of prevalence of contiguous pathways in proteins on structural complexity.PLoS One. 2017 Dec 12;12(12):e0188616. doi: 10.1371/journal.pone.0188616. eCollection 2017. PLoS One. 2017. PMID: 29232711 Free PMC article.
References
-
- Malinow R., Malenka R.C. AMPA receptor trafficking and synaptic plasticity. Annu. Rev. Neurosci. 2002;25:103–126. - PubMed
-
- Chang P.K., Verbich D., McKinney R.A. AMPA receptors as drug targets in neurological disease—advantages, caveats, and future outlook. Eur. J. Neurosci. 2012;35:1908–1916. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

