The Human Gut Microbiome as a Transporter of Antibiotic Resistance Genes between Continents
- PMID: 26259788
- PMCID: PMC4576037
- DOI: 10.1128/AAC.00933-15
The Human Gut Microbiome as a Transporter of Antibiotic Resistance Genes between Continents
Abstract
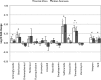
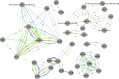
Previous studies of antibiotic resistance dissemination by travel have, by targeting only a select number of cultivable bacterial species, omitted most of the human microbiome. Here, we used explorative shotgun metagenomic sequencing to address the abundance of >300 antibiotic resistance genes in fecal specimens from 35 Swedish students taken before and after exchange programs on the Indian peninsula or in Central Africa. All specimens were additionally cultured for extended-spectrum beta-lactamase (ESBL)-producing enterobacteria, and the isolates obtained were genome sequenced. The overall taxonomic diversity and composition of the gut microbiome remained stable before and after travel, but there was an increasing abundance of Proteobacteria in 25/35 students. The relative abundance of antibiotic resistance genes increased, most prominently for genes encoding resistance to sulfonamide (2.6-fold increase), trimethoprim (7.7-fold), and beta-lactams (2.6-fold). Importantly, the increase observed occurred without any antibiotic intake. Of 18 students visiting the Indian peninsula, 12 acquired ESBL-producing Escherichia coli, while none returning from Africa were positive. Despite deep sequencing efforts, the sensitivity of metagenomics was not sufficient to detect acquisition of the low-abundant genes responsible for the observed ESBL phenotype. In conclusion, metagenomic sequencing of the intestinal microbiome of Swedish students returning from exchange programs in Central Africa or the Indian peninsula showed increased abundance of genes encoding resistance to widely used antibiotics.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures



References
-
- World Health Organization. 2014. Antimicrobial resistance: global report on surveillance. World Health Organization, Geneva, Switzerland.
-
- Ashbolt NJ, Amézquita A, Backhaus T, Borriello P, Brandt KK, Collignon P, Coors A, Finley R, Gaze WH, Heberer T, Lawrence JR, Larsson DGJ, McEwen SA, Ryan JJ, Schönfeld J, Silley P, Snape JR, Van den Eede C, Topp E. 2013. Human health risk assessment (HHRA) for environmental development and transfer of antibiotic resistance. Environ Health Perspect 121:993–1001. - PMC - PubMed
-
- Fernando GA, Collignon PJ, Bell JM. 2010. A risk for returned travellers: the “post-antibiotic era”. Med J Aust 193:59. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

