Adaptive aneuploidy protects against thiol peroxidase deficiency by increasing respiration via key mitochondrial proteins
- PMID: 26261310
- PMCID: PMC4553801
- DOI: 10.1073/pnas.1505315112
Adaptive aneuploidy protects against thiol peroxidase deficiency by increasing respiration via key mitochondrial proteins
Abstract
Aerobic respiration is a fundamental energy-generating process; however, there is cost associated with living in an oxygen-rich environment, because partially reduced oxygen species can damage cellular components. Organisms evolved enzymes that alleviate this damage and protect the intracellular milieu, most notably thiol peroxidases, which are abundant and conserved enzymes that mediate hydrogen peroxide signaling and act as the first line of defense against oxidants in nearly all living organisms. Deletion of all eight thiol peroxidase genes in yeast (∆8 strain) is not lethal, but results in slow growth and a high mutation rate. Here we characterized mechanisms that allow yeast cells to survive under conditions of thiol peroxidase deficiency. Two independent ∆8 strains increased mitochondrial content, altered mitochondrial distribution, and became dependent on respiration for growth but they were not hypersensitive to H2O2. In addition, both strains independently acquired a second copy of chromosome XI and increased expression of genes encoded by it. Survival of ∆8 cells was dependent on mitochondrial cytochrome-c peroxidase (CCP1) and UTH1, present on chromosome XI. Coexpression of these genes in ∆8 cells led to the elimination of the extra copy of chromosome XI and improved cell growth, whereas deletion of either gene was lethal. Thus, thiol peroxidase deficiency requires dosage compensation of CCP1 and UTH1 via chromosome XI aneuploidy, wherein these proteins support hydroperoxide removal with the reducing equivalents generated by the electron transport chain. To our knowledge, this is the first evidence of adaptive aneuploidy counteracting oxidative stress.
Keywords: aneuploidy; oxidative stress; respiration; thiol peroxidase.
Conflict of interest statement
The authors declare no conflict of interest.
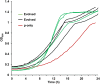
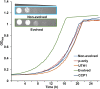
Figures






Similar articles
-
Yeast oxidative stress response. Influences of cytosolic thioredoxin peroxidase I and of the mitochondrial functional state.FEBS J. 2006 Feb;273(4):805-16. doi: 10.1111/j.1742-4658.2006.05116.x. FEBS J. 2006. PMID: 16441666
-
Thermosensitive phenotype of yeast mutant lacking thioredoxin peroxidase.Arch Biochem Biophys. 1998 Nov 1;359(1):99-106. doi: 10.1006/abbi.1998.0896. Arch Biochem Biophys. 1998. PMID: 9799566
-
Extra copy of the mitochondrial cytochrome-c peroxidase gene confers a pyruvate-underproducing characteristic of sake yeast through respiratory metabolism.J Biosci Bioeng. 2021 Jun;131(6):640-646. doi: 10.1016/j.jbiosc.2021.01.007. Epub 2021 Feb 15. J Biosci Bioeng. 2021. PMID: 33597082
-
Uth1p: a yeast mitochondrial protein at the crossroads of stress, degradation and cell death.FEMS Yeast Res. 2004 Nov;5(2):133-40. doi: 10.1016/j.femsyr.2004.05.001. FEMS Yeast Res. 2004. PMID: 15489196 Review.
-
Reduction of hydrogen peroxide in gram-negative bacteria - bacterial peroxidases.Adv Microb Physiol. 2019;74:415-464. doi: 10.1016/bs.ampbs.2019.02.006. Epub 2019 Apr 8. Adv Microb Physiol. 2019. PMID: 31126534 Review.
Cited by
-
Aneuploidy: Cancer strength or vulnerability?Int J Cancer. 2019 Jan 1;144(1):8-25. doi: 10.1002/ijc.31718. Epub 2018 Oct 31. Int J Cancer. 2019. PMID: 29981145 Free PMC article. Review.
-
Aneuploidy and gene expression: is there dosage compensation?Epigenomics. 2019 Dec;11(16):1827-1837. doi: 10.2217/epi-2019-0135. Epub 2019 Nov 22. Epigenomics. 2019. PMID: 31755744 Free PMC article. Review.
-
Thiol Peroxidases as Major Regulators of Intracellular Levels of Peroxynitrite in Live Saccharomyces cerevisiae Cells.Antioxidants (Basel). 2020 May 16;9(5):434. doi: 10.3390/antiox9050434. Antioxidants (Basel). 2020. PMID: 32429358 Free PMC article.
-
Systematic analysis of bypass suppression of essential genes.Mol Syst Biol. 2020 Sep;16(9):e9828. doi: 10.15252/msb.20209828. Mol Syst Biol. 2020. PMID: 32939983 Free PMC article.
-
Aneuploidy of specific chromosomes is beneficial to cells lacking spindle checkpoint protein Bub3.PLoS Genet. 2025 Feb 4;21(2):e1011576. doi: 10.1371/journal.pgen.1011576. eCollection 2025 Feb. PLoS Genet. 2025. PMID: 39903784 Free PMC article.
References
-
- Apel K, Hirt H. Reactive oxygen species: Metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol. 2004;55:373–399. - PubMed
-
- Cross CE, et al. Oxygen radicals and human disease. Ann Intern Med. 1987;107(4):526–545. - PubMed
-
- Rhee SG, Chae HZ, Kim K. Peroxiredoxins: A historical overview and speculative preview of novel mechanisms and emerging concepts in cell signaling. Free Radic Biol Med. 2005;38(12):1543–1552. - PubMed
Publication types
MeSH terms
Substances
Associated data
- BioProject/PRJNA287442
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

