New paradigms in GPCR drug discovery
- PMID: 26265138
- PMCID: PMC4967540
- DOI: 10.1016/j.bcp.2015.08.085
New paradigms in GPCR drug discovery
Abstract
G protein-coupled receptors (GPCRs) remain a major domain of pharmaceutical discovery. The identification of GPCR lead compounds and their optimization are now structure-based, thanks to advances in X-ray crystallography, molecular modeling, protein engineering and biophysical techniques. In silico screening provides useful hit molecules. New pharmacological approaches to tuning the pleotropic action of GPCRs include: allosteric modulators, biased ligands, GPCR heterodimer-targeted compounds, manipulation of polypharmacology, receptor antibodies and tailoring of drug molecules to fit GPCR pharmacogenomics. Measurements of kinetics and drug efficacy are factors influencing clinical success. With the exception of inhibitors of GPCR kinases, targeting of intracellular GPCR signaling or receptor cycling for therapeutic purposes remains a futuristic concept. New assay approaches are more efficient and multidimensional: cell-based, label-free, fluorescence-based assays, and biosensors. Tailoring GPCR drugs to a patient's genetic background is now being considered. Chemoinformatic tools can predict ADME-tox properties. New imaging technology visualizes drug action in vivo. Thus, there is reason to be optimistic that new technology for GPCR ligand discovery will help reverse the current narrowing of the pharmaceutical pipeline.
Keywords: Drug discovery; GPCR; Inhibitors; Signaling; X-ray crystallography; structure-based design.
Published by Elsevier Inc.
Figures

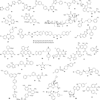
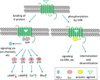
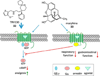






References
-
- Beitelshees AL, McLeod HL. Clopidogrel pharmacogenetics promising steps towards patient care? Arteriosclerosis, Thrombosis, and Vascular Biology. 2006;26:1681–1683. - PubMed
-
- Zhang X, Wang H, Ma Z, Wu B. Effects of pharmaceutical PEGylation on drug metabolism and its clinical concerns. Expert Opin Drug Metab Toxicol. 2014;10:1691–1702. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

