Achieving high-sensitivity for clinical applications using augmented exome sequencing
- PMID: 26269718
- PMCID: PMC4534066
- DOI: 10.1186/s13073-015-0197-4
Achieving high-sensitivity for clinical applications using augmented exome sequencing
Abstract
Background: Whole exome sequencing is increasingly used for the clinical evaluation of genetic disease, yet the variation of coverage and sensitivity over medically relevant parts of the genome remains poorly understood. Several sequencing-based assays continue to provide coverage that is inadequate for clinical assessment.
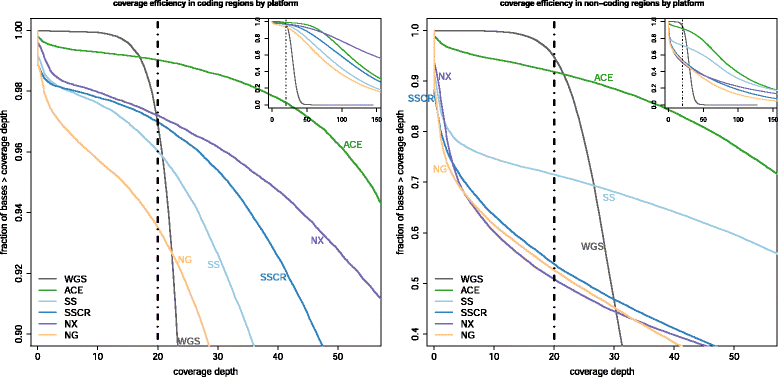
Methods: Using sequence data obtained from the NA12878 reference sample and pre-defined lists of medically-relevant protein-coding and noncoding sequences, we compared the breadth and depth of coverage obtained among four commercial exome capture platforms and whole genome sequencing. In addition, we evaluated the performance of an augmented exome strategy, ACE, that extends coverage in medically relevant regions and enhances coverage in areas that are challenging to sequence. Leveraging reference call-sets, we also examined the effects of improved coverage on variant detection sensitivity.
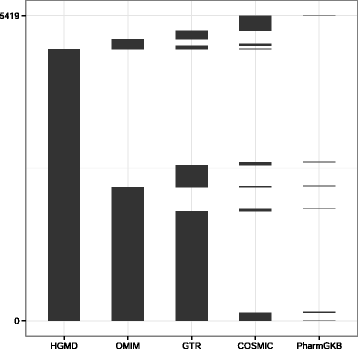
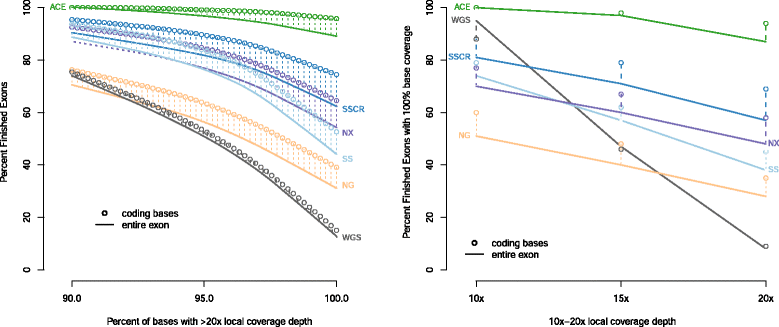
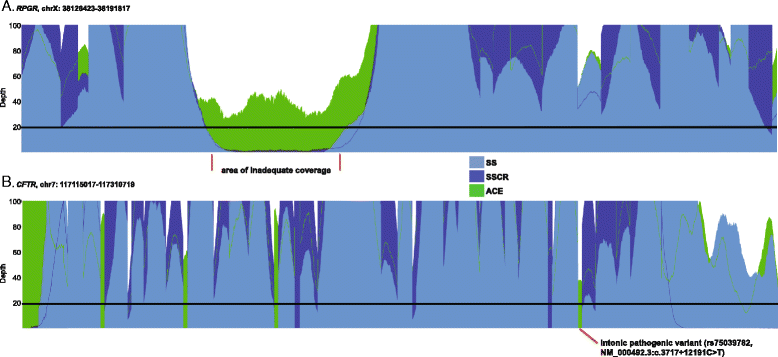
Results: We observed coverage shortfalls with each of the conventional exome-capture and whole-genome platforms across several medically interpretable genes. These gaps included areas of the genome required for reporting recently established secondary findings (ACMG) and known disease-associated loci. The augmented exome strategy recovered many of these gaps, resulting in improved coverage in these areas. At clinically-relevant coverage levels (100 % bases covered at ≥20×), ACE improved coverage among genes in the medically interpretable genome (>90 % covered relative to 10-78 % with other platforms), the set of ACMG secondary finding genes (91 % covered relative to 4-75 % with other platforms) and a subset of variants known to be associated with human disease (99 % covered relative to 52-95 % with other platforms). Improved coverage translated into improvements in sensitivity, with ACE variant detection sensitivities (>97.5 % SNVs, >92.5 % InDels) exceeding that observed with conventional whole-exome and whole-genome platforms.
Conclusions: Clinicians should consider analytical performance when making clinical assessments, given that even a few missed variants can lead to reporting false negative results. An augmented exome strategy provides a level of coverage not achievable with other platforms, thus addressing concerns regarding the lack of sensitivity in clinically important regions. In clinical applications where comprehensive coverage of medically interpretable areas of the genome requires higher localized sequencing depth, an augmented exome approach offers both cost and performance advantages over other sequencing-based tests.
Figures


References
-
- Gahl WA, Markello TC, Toro C, Fajardo KF, Sincan M, Gill F, Carlson-Donohoe H, Gropman A, Pierson TM, Golas G, Wolfe L, Groden C, Godfrey R, Nehrebecky M, Wahl C, Landis DMD, Yang S, Madeo A, Mullikin JC, Boerkoel CF, Tifft CJ, Adams D. The National Institutes of Health Undiagnosed Diseases Program: insights into rare diseases. Genet Med. 2012;14:51–9. doi: 10.1038/gim.0b013e318232a005. - DOI - PMC - PubMed
-
- Lupski JR, Gonzaga-Jauregui C, Yang Y, Bainbridge MN, Jhangiani S, Buhay CJ, Kovar CL, Wang M, Hawes AC, Reid JG, Eng C, Muzny DM, Gibbs RA. Exome sequencing resolves apparent incidental findings and reveals further complexity of SH3TC2 variant alleles causing Charcot-Marie-Tooth neuropathy. Genome Med. 2013;5:57. doi: 10.1186/gm461. - DOI - PMC - PubMed
-
- Yang Y, Muzny DM, Reid JG, Bainbridge MN, Willis A, Ward PA, Braxton A, Beuten J, Xia F, Niu Z, Hardison M, Person R, Bekheirnia MR, Leduc MS, Kirby A, Pham P, Scull J, Wang M, Ding Y, Plon SE, Lupski JR, Beaudet AL, Gibbs RA, Eng CM. Clinical whole-exome sequencing for the diagnosis of mendelian disorders. N Engl J Med. 2013;369:1502–11. doi: 10.1056/NEJMoa1306555. - DOI - PMC - PubMed
-
- Neveling K, Feenstra I, Gilissen C, Hoefsloot LH, Kamsteeg E-J, Mensenkamp AR, Rodenburg RJT, Yntema HG, Spruijt L, Vermeer S, Rinne T, van Gassen KL, Bodmer D, Lugtenberg D, de Reuver R, Buijsman W, Derks RC, Wieskamp N, van den Heuvel B, Ligtenberg MJL, Kremer H, Koolen DA, van de Warrenburg BPC, Cremers FPM, Marcelis CLM, Smeitink JAM, Wortmann SB, van Zelst-Stams WAG, Veltman JA, Brunner HG, et al. A post-hoc comparison of the utility of sanger sequencing and exome sequencing for the diagnosis of heterogeneous diseases. Hum Mutat. 2013;34:1721–6. doi: 10.1002/humu.22450. - DOI - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

