mTOR regulates MAPKAPK2 translation to control the senescence-associated secretory phenotype
- PMID: 26280535
- PMCID: PMC4589897
- DOI: 10.1038/ncb3225
mTOR regulates MAPKAPK2 translation to control the senescence-associated secretory phenotype
Erratum in
-
Erratum: mTOR regulates MAPKAPK2 translation to control the senescence-associated secretory phenotype.Nat Cell Biol. 2015 Oct;17(10):1370. doi: 10.1038/ncb3243. Nat Cell Biol. 2015. PMID: 26419804 No abstract available.
-
Author Correction: mTOR regulates MAPKAPK2 translation to control the senescence-associated secretory phenotype.Nat Cell Biol. 2024 Jun;26(6):1019. doi: 10.1038/s41556-024-01443-6. Nat Cell Biol. 2024. PMID: 38778130 No abstract available.
Abstract
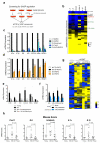
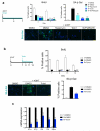
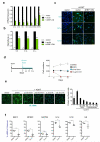
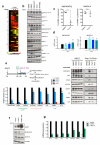
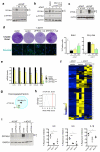
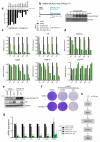
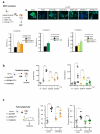
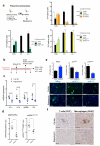
Senescent cells secrete a combination of factors collectively known as the senescence-associated secretory phenotype (SASP). The SASP reinforces senescence and activates an immune surveillance response, but it can also show pro-tumorigenic properties and contribute to age-related pathologies. In a drug screen to find new SASP regulators, we uncovered the mTOR inhibitor rapamycin as a potent SASP suppressor. Here we report a mechanism by which mTOR controls the SASP by differentially regulating the translation of the MK2 (also known as MAPKAPK2) kinase through 4EBP1. In turn, MAPKAPK2 phosphorylates the RNA-binding protein ZFP36L1 during senescence, inhibiting its ability to degrade the transcripts of numerous SASP components. Consequently, mTOR inhibition or constitutive activation of ZFP36L1 impairs the non-cell-autonomous effects of senescent cells in both tumour-suppressive and tumour-promoting contexts. Altogether, our results place regulation of the SASP as a key mechanism by which mTOR could influence cancer, age-related diseases and immune responses.
Figures
Comment in
-
The InflammTORy Powers of Senescence.Trends Cell Biol. 2015 Nov;25(11):634-636. doi: 10.1016/j.tcb.2015.09.011. Epub 2015 Oct 21. Trends Cell Biol. 2015. PMID: 26471225
References
METHODS REFERENCES
-
- Fellmann C, et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 2013;5:1704–1713. - PubMed
-
- Debacq-Chainiaux F, Erusalimsky JD, Campisi J, Toussaint O. Protocols to detect senescence-associated beta-galactosidase (SA-betagal) activity, a biomarker of senescent cells in culture and in vivo. Nat Protoc. 2009;4:1798–1806. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

