Diagnostic interpretation of array data using public databases and internet sources
- PMID: 26285306
- PMCID: PMC5027376
- DOI: 10.1002/humu.22049
Diagnostic interpretation of array data using public databases and internet sources
Abstract
The range of commercially available array platforms and analysis software packages is expanding and their utility is improving, making reliable detection of copy-number variants (CNVs) relatively straightforward. Reliable interpretation of CNV data, however, is often difficult and requires expertise. With our knowledge of the human genome growing rapidly, applications for array testing continuously broadening, and the resolution of CNV detection increasing, this leads to great complexity in interpreting what can be daunting data. Correct CNV interpretation and optimal use of the genotype information provided by single-nucleotide polymorphism probes on an array depends largely on knowledge present in various resources. In addition to the availability of host laboratories' own datasets and national registries, there are several public databases and Internet resources with genotype and phenotype information that can be used for array data interpretation. With so many resources now available, it is important to know which are fit-for-purpose in a diagnostic setting. We summarize the characteristics of the most commonly used Internet databases and resources, and propose a general data interpretation strategy that can be used for comparative hybridization, comparative intensity, and genotype-based array data.
Conflict of interest statement
Statement: The authors declare no conflict of interest.
Figures
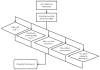
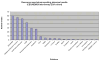

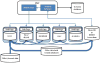
Similar articles
-
CNV Workshop: an integrated platform for high-throughput copy number variation discovery and clinical diagnostics.BMC Bioinformatics. 2010 Feb 4;11:74. doi: 10.1186/1471-2105-11-74. BMC Bioinformatics. 2010. PMID: 20132550 Free PMC article.
-
Comprehensive performance comparison of high-resolution array platforms for genome-wide Copy Number Variation (CNV) analysis in humans.BMC Genomics. 2017 Apr 24;18(1):321. doi: 10.1186/s12864-017-3658-x. BMC Genomics. 2017. PMID: 28438122 Free PMC article.
-
CNVinspector: a web-based tool for the interactive evaluation of copy number variations in single patients and in cohorts.J Med Genet. 2013 Aug;50(8):529-33. doi: 10.1136/jmedgenet-2012-101497. Epub 2013 May 31. J Med Genet. 2013. PMID: 23729504
-
Detection and interpretation of genomic structural variation in health and disease.Expert Rev Mol Diagn. 2013 Jan;13(1):61-82. doi: 10.1586/erm.12.119. Expert Rev Mol Diagn. 2013. PMID: 23256704 Review.
-
SNP array analysis in constitutional and cancer genome diagnostics--copy number variants, genotyping and quality control.Cytogenet Genome Res. 2011;135(3-4):212-21. doi: 10.1159/000331273. Epub 2011 Sep 16. Cytogenet Genome Res. 2011. PMID: 21934286 Review.
Cited by
-
European guidelines for constitutional cytogenomic analysis.Eur J Hum Genet. 2019 Jan;27(1):1-16. doi: 10.1038/s41431-018-0244-x. Epub 2018 Oct 1. Eur J Hum Genet. 2019. PMID: 30275486 Free PMC article. Review.
-
Dynamic Interplay between Structural Variations and 3D Genome Organization in Pancreatic Cancer.Adv Sci (Weinh). 2022 Jun;9(18):e2200818. doi: 10.1002/advs.202200818. Epub 2022 May 15. Adv Sci (Weinh). 2022. PMID: 35570408 Free PMC article.
-
Deciphering the pathogenic consequences of chromosomal aberrations in human genetic disease.Mol Cytogenet. 2014 Dec 19;7(1):100. doi: 10.1186/s13039-014-0100-9. eCollection 2014. Mol Cytogenet. 2014. PMID: 25606056 Free PMC article.
-
Identification of Key Genes and Pathways in Post-traumatic Stress Disorder Using Microarray Analysis.Front Psychol. 2019 Feb 27;10:302. doi: 10.3389/fpsyg.2019.00302. eCollection 2019. Front Psychol. 2019. PMID: 30873067 Free PMC article.
-
Identification of key genes in human airway epithelial cells in response to respiratory pathogens using microarray analysis.BMC Microbiol. 2018 Jun 8;18(1):58. doi: 10.1186/s12866-018-1187-7. BMC Microbiol. 2018. PMID: 29884128 Free PMC article.
References
-
- Conrad DF, Pinto D, Redon R, Feuk L, Gokcumen O, Zhang Y, Aerts J, Andrews TD, Barnes C, Campbell P, Fitzgerald T, Hu M, Ihm CH, Kristiansson K, Macarthur DG, Macdonald JR, Onyiah I, Pang AW, Robson S, Stirrups K, Valsesia A, Walter K, Wei J, Tyler-Smith C, Carter NP, Lee C, Scherer SW, Hurles ME Wellcome Trust Case Control Consortium. Origins and functional impact of copy number variation in the human genome. Nature. 2010;464:704–412. - PMC - PubMed
-
- Demars J, Rossignol S, Netchine I, Lee KS, Shmela M, Faivre L, Weill J, Odent S, Azzi S, Callier P, Lucas J, Dubourg C, Andrieux J, Bouc YL, El-Osta A, Gicquel C. New insights into the pathogenesis of Beckwith-Wiedemann and Silver-Russell syndromes: contribution of small copy number variations to 11p15 imprinting defects. Hum Mutat. 2011;32:1171–1182. - PubMed
-
- Feenstra I, Fang J, Koolen DA, Siezen A, Evans C, Winter RM, Lees MM, Riegel M, de Vries BBA, van Ravenswaaij CMA, Schinzel A. European cytogenetics association register of unbalanced chromosome aberrations (ECARUCA): an online database for rare chromosomal abnormalities. Eur J Med Genet. 2006;49:279–291. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

