Limiting replication stress during somatic cell reprogramming reduces genomic instability in induced pluripotent stem cells
- PMID: 26292731
- PMCID: PMC4560784
- DOI: 10.1038/ncomms9036
Limiting replication stress during somatic cell reprogramming reduces genomic instability in induced pluripotent stem cells
Abstract
The generation of induced pluripotent stem cells (iPSC) from adult somatic cells is one of the most remarkable discoveries in recent decades. However, several works have reported evidence of genomic instability in iPSC, raising concerns on their biomedical use. The reasons behind the genomic instability observed in iPSC remain mostly unknown. Here we show that, similar to the phenomenon of oncogene-induced replication stress, the expression of reprogramming factors induces replication stress. Increasing the levels of the checkpoint kinase 1 (CHK1) reduces reprogramming-induced replication stress and increases the efficiency of iPSC generation. Similarly, nucleoside supplementation during reprogramming reduces the load of DNA damage and genomic rearrangements on iPSC. Our data reveal that lowering replication stress during reprogramming, genetically or chemically, provides a simple strategy to reduce genomic instability on mouse and human iPSC.
Figures
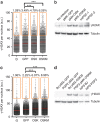

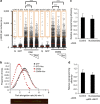
 ) were also included as negative control. Data are representative of three independent experiments performed in triplicate ***P<0.001. (b) Graphical representation of the fork elongation rate (kb min−1), measured by DNA combing, in BJ fibroblasts 4 days after infection with retroviruses encoding either GFP or OSKM with or without the addition of nucleosides. A representative image of a stained DNA fibre is shown in the lower part. Iododeoxyuridine (IdU, red) and Chlorodeoxyuridine (CIdU, green) signals are shown. (c,d) Relative reprogramming efficiency determined in RFP-infected (c) or RFP plus cMyc-infected (d) dFib-indOSK cells treated with doxycycline at a concentration of 100 ng ml−1 with or without daily addition of nucleosides. Reprogramming efficiency was evaluated by scoring the percentage of GFP-positive colonies (resulting from the reactivation of the endogenous OCT4 promoter) and RFP-negative colonies (resulting from the silencing of exogenous transgenes). Data are representative of two independent experiments performed in duplicate.
) were also included as negative control. Data are representative of three independent experiments performed in triplicate ***P<0.001. (b) Graphical representation of the fork elongation rate (kb min−1), measured by DNA combing, in BJ fibroblasts 4 days after infection with retroviruses encoding either GFP or OSKM with or without the addition of nucleosides. A representative image of a stained DNA fibre is shown in the lower part. Iododeoxyuridine (IdU, red) and Chlorodeoxyuridine (CIdU, green) signals are shown. (c,d) Relative reprogramming efficiency determined in RFP-infected (c) or RFP plus cMyc-infected (d) dFib-indOSK cells treated with doxycycline at a concentration of 100 ng ml−1 with or without daily addition of nucleosides. Reprogramming efficiency was evaluated by scoring the percentage of GFP-positive colonies (resulting from the reactivation of the endogenous OCT4 promoter) and RFP-negative colonies (resulting from the silencing of exogenous transgenes). Data are representative of two independent experiments performed in duplicate.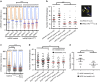
References
-
- Takahashi K. & Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006). - PubMed
-
- Hussein S. M. et al. Copy number variation and selection during reprogramming to pluripotency. Nature 471, 58–62 (2011). - PubMed
-
- Ji J. et al. Elevated coding mutation rate during the reprogramming of human somatic cells into induced pluripotent stem cells. Stem Cells 30, 435–440 (2012). - PubMed
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

