Protein folding guides disulfide bond formation
- PMID: 26297249
- PMCID: PMC4568676
- DOI: 10.1073/pnas.1503909112
Protein folding guides disulfide bond formation
Abstract
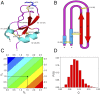
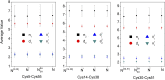
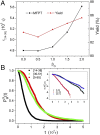
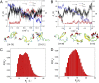
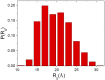
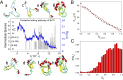
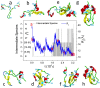
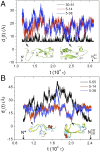
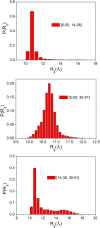
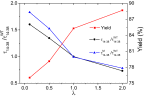
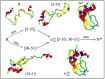
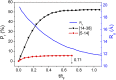
The Anfinsen principle that the protein sequence uniquely determines its structure is based on experiments on oxidative refolding of a protein with disulfide bonds. The problem of how protein folding drives disulfide bond formation is poorly understood. Here, we have solved this long-standing problem by creating a general method for implementing the chemistry of disulfide bond formation and rupture in coarse-grained molecular simulations. As a case study, we investigate the oxidative folding of bovine pancreatic trypsin inhibitor (BPTI). After confirming the experimental findings that the multiple routes to the folded state contain a network of states dominated by native disulfides, we show that the entropically unfavorable native single disulfide [14-38] between Cys14 and Cys38 forms only after polypeptide chain collapse and complete structuring of the central core of the protein containing an antiparallel β-sheet. Subsequent assembly, resulting in native two-disulfide bonds and the folded state, involves substantial unfolding of the protein and transient population of nonnative structures. The rate of [14-38] formation increases as the β-sheet stability increases. The flux to the native state, through a network of kinetically connected native-like intermediates, changes dramatically by altering the redox conditions. Disulfide bond formation between Cys residues not present in the native state are relevant only on the time scale of collapse of BPTI. The finding that formation of specific collapsed native-like structures guides efficient folding is applicable to a broad class of single-domain proteins, including enzyme-catalyzed disulfide proteins.
Keywords: disulfide proteins; early collapse; enzyme-catalyzed folding; native-like interactions; nonnative interactions.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Anfinsen CB. Principles that govern the folding of protein chains. Science. 1973;181(4096):223–230. - PubMed
-
- Anfinsen CB, Scheraga HA. Experimental and theoretical aspects of protein folding. Adv Protein Chem. 1975;29:205–300. - PubMed
-
- Schröder M, Kaufman RJ. The mammalian unfolded protein response. Annu Rev Biochem. 2005;74:739–789. - PubMed
-
- Welker E, Wedemeyer WJ, Narayan M, Scheraga HA. Coupling of conformational folding and disulfide-bond reactions in oxidative folding of proteins. Biochemistry. 2001;40(31):9059–9064. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

