Global Promoter Targeting of a Conserved Lysine Deacetylase for Transcriptional Shutoff during Quiescence Entry
- PMID: 26300265
- PMCID: PMC4560983
- DOI: 10.1016/j.molcel.2015.07.014
Global Promoter Targeting of a Conserved Lysine Deacetylase for Transcriptional Shutoff during Quiescence Entry
Abstract
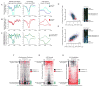
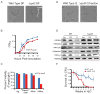
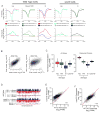
Quiescence is a conserved cell-cycle state characterized by cell-cycle arrest, increased stress resistance, enhanced longevity, and decreased transcriptional, translational, and metabolic output. Although quiescence plays essential roles in cell survival and normal differentiation, the molecular mechanisms leading to this state are not well understood. Here, we determined changes in the transcriptome and chromatin structure of S. cerevisiae upon quiescence entry. Our analyses revealed transcriptional shutoff that is far more robust than previously believed and an unprecedented global chromatin transition, which are tightly correlated. These changes require Rpd3 lysine deacetylase targeting to at least half of gene promoters via quiescence-specific transcription factors including Xbp1 and Stb3. Deletion of RPD3 prevents cells from establishing transcriptional quiescence, leading to defects in quiescence entry and shortening of chronological lifespan. Our results define a molecular mechanism for global reprogramming of transcriptome and chromatin structure for quiescence driven by a highly conserved chromatin regulator.
Copyright © 2015 Elsevier Inc. All rights reserved.
Figures



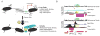
References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

