Partially Unfolded Forms of the Prion Protein Populated under Misfolding-promoting Conditions: CHARACTERIZATION BY HYDROGEN EXCHANGE MASS SPECTROMETRY AND NMR
- PMID: 26306043
- PMCID: PMC4646174
- DOI: 10.1074/jbc.M115.677575
Partially Unfolded Forms of the Prion Protein Populated under Misfolding-promoting Conditions: CHARACTERIZATION BY HYDROGEN EXCHANGE MASS SPECTROMETRY AND NMR
Abstract
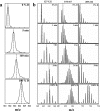
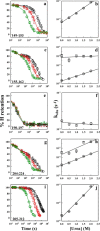
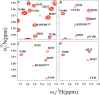
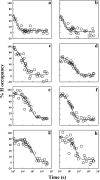
The susceptibility of the cellular prion protein (PrP(C)) to convert to an alternative misfolded conformation (PrP(Sc)), which is the key event in the pathogenesis of prion diseases, is indicative of a conformationally flexible native (N) state. In the present study, hydrogen-deuterium exchange (HDX) in conjunction with mass spectrometry and nuclear magnetic resonance spectroscopy were used for the structural and energetic characterization of the N state of the full-length mouse prion protein, moPrP(23-231), under conditions that favor misfolding. The kinetics of HDX of 34 backbone amide hydrogens in the N state were determined at pH 4. In contrast to the results of previous HDX studies on the human and Syrian hamster prion proteins at a higher pH, various segments of moPrP were found to undergo different extents of subglobal unfolding events at pH 4, a pH at which the protein is known to be primed to misfold to a β-rich conformation. No residual structure around the disulfide bond was observed for the unfolded state at pH 4. The N state of the prion protein was observed to be at equilibrium with at least two partially unfolded forms (PUFs). These PUFs, which are accessed by stochastic fluctuations of the N state, have altered surface area exposure relative to the N state. One of these PUFs resembles a conformation previously implicated to be an initial intermediate in the conversion of monomeric protein into misfolded oligomer at pH 4.
Keywords: hydrogen-deuterium exchange; mass spectrometry (MS); nuclear magnetic resonance (NMR); partially unfolded forms; prion; protein dynamic.
© 2015 by The American Society for Biochemistry and Molecular Biology, Inc.
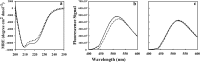
Figures




Similar articles
-
Identification and Structural Characterization of the Precursor Conformation of the Prion Protein which Directly Initiates Misfolding and Oligomerization.J Mol Biol. 2017 Mar 24;429(6):886-899. doi: 10.1016/j.jmb.2017.01.019. Epub 2017 Jan 29. J Mol Biol. 2017. PMID: 28147229
-
Mutations of evolutionarily conserved aromatic residues suggest that misfolding of the mouse prion protein may commence in multiple ways.J Neurochem. 2023 Dec;167(5):696-710. doi: 10.1111/jnc.16007. Epub 2023 Nov 8. J Neurochem. 2023. PMID: 37941487
-
Slow Misfolding of a Molten Globule form of a Mutant Prion Protein Variant into a β-rich Dimer.J Mol Biol. 2024 Oct 1;436(19):168736. doi: 10.1016/j.jmb.2024.168736. Epub 2024 Aug 5. J Mol Biol. 2024. PMID: 39097185
-
Prion protein oligomer and its neurotoxicity.Acta Biochim Biophys Sin (Shanghai). 2013 Jun;45(6):442-51. doi: 10.1093/abbs/gmt037. Epub 2013 Apr 4. Acta Biochim Biophys Sin (Shanghai). 2013. PMID: 23557632 Review.
-
Measuring the hydrogen/deuterium exchange of proteins at high spatial resolution by mass spectrometry: overcoming gas-phase hydrogen/deuterium scrambling.Acc Chem Res. 2014 Oct 21;47(10):3018-27. doi: 10.1021/ar500194w. Epub 2014 Aug 29. Acc Chem Res. 2014. PMID: 25171396 Review.
Cited by
-
The Pathogenic A116V Mutation Enhances Ion-Selective Channel Formation by Prion Protein in Membranes.Biophys J. 2016 Apr 26;110(8):1766-1776. doi: 10.1016/j.bpj.2016.03.017. Biophys J. 2016. Retraction in: Biophys J. 2021 Dec 7;120(23):5421. doi: 10.1016/j.bpj.2021.11.005. PMID: 27119637 Free PMC article. Retracted.
-
Identification of Two Early Folding Stage Prion Non-Local Contacts Suggested to Serve as Key Steps in Directing the Final Fold to Be Either Native or Pathogenic.Int J Mol Sci. 2021 Aug 10;22(16):8619. doi: 10.3390/ijms22168619. Int J Mol Sci. 2021. PMID: 34445324 Free PMC article.
-
Structural and dynamical determinants of a β-sheet-enriched intermediate involved in amyloid fibrillar assembly of human prion protein.Chem Sci. 2022 Aug 5;13(35):10406-10427. doi: 10.1039/d2sc00345g. eCollection 2022 Sep 14. Chem Sci. 2022. PMID: 36277622 Free PMC article.
-
Conformational Enigma of TDP-43 Misfolding in Neurodegenerative Disorders.ACS Omega. 2024 Sep 20;9(39):40286-40297. doi: 10.1021/acsomega.4c04119. eCollection 2024 Oct 1. ACS Omega. 2024. PMID: 39372031 Free PMC article. Review.
-
Destabilization of polar interactions in the prion protein triggers misfolding and oligomerization.Protein Sci. 2021 Nov;30(11):2258-2271. doi: 10.1002/pro.4188. Epub 2021 Sep 30. Protein Sci. 2021. PMID: 34558139 Free PMC article.
References
-
- Gajdusek D. C. (1977) Unconventional viruses and the origin and disappearance of kuru. Science 197, 943–960 - PubMed
-
- Prusiner S. B. (1982) Novel proteinaceous infectious particles cause scrapie. Science 216, 136–144 - PubMed
-
- Prusiner S. B. (1991) Molecular biology of prion diseases. Science 252, 1515–1522 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Research Materials

