Novel roles for LIX1L in promoting cancer cell proliferation through ROS1-mediated LIX1L phosphorylation
- PMID: 26310847
- PMCID: PMC4550850
- DOI: 10.1038/srep13474
Novel roles for LIX1L in promoting cancer cell proliferation through ROS1-mediated LIX1L phosphorylation
Abstract
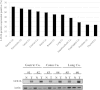
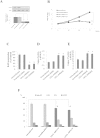
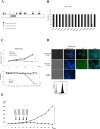
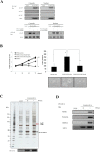
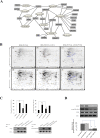
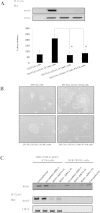
Herein, we report the characterization of Limb expression 1-like, (LIX1L), a putative RNA-binding protein (RBP) containing a double-stranded RNA binding motif, which is highly expressed in various cancer tissues. Analysis of MALDI-TOF/TOF mass spectrometry and RNA immunoprecipitation-sequencing of interacting proteins and the microRNAs (miRNAs) bound to LIX1L revealed that LIX1L interacts with proteins (RIOK1, nucleolin and PABPC4) and miRNAs (has-miRNA-520a-5p, -300, -216b, -326, -190a, -548b-3p, -7-5p and -1296) in HEK-293 cells. Moreover, the reduction of phosphorylated Tyr(136) (pTyr(136)) in LIX1L through the homeodomain peptide, PY136, inhibited LIX1L-induced cell proliferation in vitro, and PY136 inhibited MKN45 cell proliferation in vivo. We also determined the miRNA-targeted genes and showed that was apoptosis induced through the reduction of pTyr(136). Moreover, ROS1, HCK, ABL1, ABL2, JAK3, LCK and TYR03 were identified as candidate kinases responsible for the phosphorylation of Tyr(136) of LIX1L. These data provide novel insights into the biological significance of LIX1L, suggesting that this protein might be an RBP, with implications for therapeutic approaches for targeting LIX1L in LIX1L-expressing cancer cells.
Figures
References
-
- Swindell E. C., Moeller C., Thaller C. & Eichele G. Cloning and expression analysis of chicken Lix1, a founding member of a novel gene family. Mech Dev. 109, 405–408 (2001). - PubMed
-
- Moeller C., Yaylaoglu M. B., Alvarez-Bolado G., Thaller C. & Eichele G. Murine Lix1, a novel marker for substantia nigra, cortical layer 5, and hindbrain structures. Brain Res Gene Expr Patterns. 1, 199–203 (2002). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

