Parvalbumin+ Neurons and Npas1+ Neurons Are Distinct Neuron Classes in the Mouse External Globus Pallidus
- PMID: 26311767
- PMCID: PMC4549397
- DOI: 10.1523/JNEUROSCI.4672-14.2015
Parvalbumin+ Neurons and Npas1+ Neurons Are Distinct Neuron Classes in the Mouse External Globus Pallidus
Abstract
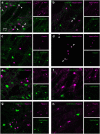
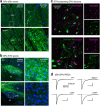
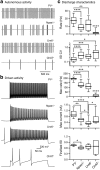
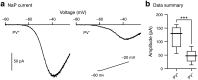
Compelling evidence suggests that pathological activity of the external globus pallidus (GPe), a nucleus in the basal ganglia, contributes to the motor symptoms of a variety of movement disorders such as Parkinson's disease. Recent studies have challenged the idea that the GPe comprises a single, homogenous population of neurons that serves as a simple relay in the indirect pathway. However, we still lack a full understanding of the diversity of the neurons that make up the GPe. Specifically, a more precise classification scheme is needed to better describe the fundamental biology and function of different GPe neuron classes. To this end, we generated a novel multicistronic BAC (bacterial artificial chromosome) transgenic mouse line under the regulatory elements of the Npas1 gene. Using a combinatorial transgenic and immunohistochemical approach, we discovered that parvalbumin-expressing neurons and Npas1-expressing neurons in the GPe represent two nonoverlapping cell classes, amounting to 55% and 27% of the total GPe neuron population, respectively. These two genetically identified cell classes projected primarily to the subthalamic nucleus and to the striatum, respectively. Additionally, parvalbumin-expressing neurons and Npas1-expressing neurons were distinct in their autonomous and driven firing characteristics, their expression of intrinsic ion conductances, and their responsiveness to chronic 6-hydroxydopamine lesion. In summary, our data argue that parvalbumin-expressing neurons and Npas1-expressing neurons are two distinct functional classes of GPe neurons. This work revises our understanding of the GPe, and provides the foundation for future studies of its function and dysfunction.
Significance statement: Until recently, the heterogeneity of the constituent neurons within the external globus pallidus (GPe) was not fully appreciated. We addressed this knowledge gap by discovering two principal GPe neuron classes, which were identified by their nonoverlapping expression of the markers parvalbumin and Npas1. Our study provides evidence that parvalbumin and Npas1 neurons have different topologies within the basal ganglia.
Keywords: 6-OHDA; BAC transgenic mice; HCN; Kv4; Nav; intrinsic properties.
Copyright © 2015 the authors 0270-6474/15/3511830-18$15.00/0.
Figures
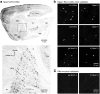
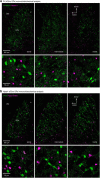
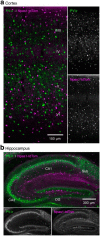
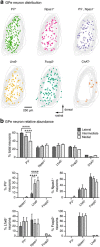


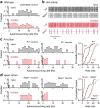

References
-
- Abdi A, Mallet N, Mohamed FY, Sharott A, Dodson PD, Nakamura KC, Suri S, Avery SV, Larvin JT, Garas FN, Garas SN, Vinciati F, Morin S, Bezard E, Baufreton J, Magill PJ. Prototypic and arkypallidal neurons in the dopamine-intact external globus pallidus. J Neurosci. 2015;35:6667–6688. doi: 10.1523/JNEUROSCI.4662-14.2015. - DOI - PMC - PubMed
-
- Anderson ME, Horak FB. Influence of the globus pallidus on arm movements in monkeys. III. Timing of movement-related information. J Neurophysiol. 1985;54:433–448. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- AG-020506/AG/NIA NIH HHS/United States
- K01 AG036738/AG/NIA NIH HHS/United States
- T32 NS041234/NS/NINDS NIH HHS/United States
- R01 NS069777/NS/NINDS NIH HHS/United States
- NS-069777-S1/NS/NINDS NIH HHS/United States
- NS-047085/NS/NINDS NIH HHS/United States
- P50 NS047085/NS/NINDS NIH HHS/United States
- NS-041234/NS/NINDS NIH HHS/United States
- F32 AG047782/AG/NIA NIH HHS/United States
- AG-047782/AG/NIA NIH HHS/United States
- T32 AG020506/AG/NIA NIH HHS/United States
- NS-069777/NS/NINDS NIH HHS/United States
- R56 NS069777/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
