Escher: A Web Application for Building, Sharing, and Embedding Data-Rich Visualizations of Biological Pathways
- PMID: 26313928
- PMCID: PMC4552468
- DOI: 10.1371/journal.pcbi.1004321
Escher: A Web Application for Building, Sharing, and Embedding Data-Rich Visualizations of Biological Pathways
Abstract
Escher is a web application for visualizing data on biological pathways. Three key features make Escher a uniquely effective tool for pathway visualization. First, users can rapidly design new pathway maps. Escher provides pathway suggestions based on user data and genome-scale models, so users can draw pathways in a semi-automated way. Second, users can visualize data related to genes or proteins on the associated reactions and pathways, using rules that define which enzymes catalyze each reaction. Thus, users can identify trends in common genomic data types (e.g. RNA-Seq, proteomics, ChIP)--in conjunction with metabolite- and reaction-oriented data types (e.g. metabolomics, fluxomics). Third, Escher harnesses the strengths of web technologies (SVG, D3, developer tools) so that visualizations can be rapidly adapted, extended, shared, and embedded. This paper provides examples of each of these features and explains how the development approach used for Escher can be used to guide the development of future visualization tools.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

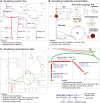


References
-
- Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: A web-based environment for protein structure homology modelling. Bioinformatics 22: 195–201. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

