Genomic expression program of Saccharomyces cerevisiae along a mixed-culture wine fermentation with Hanseniaspora guilliermondii
- PMID: 26314747
- PMCID: PMC4552253
- DOI: 10.1186/s12934-015-0318-1
Genomic expression program of Saccharomyces cerevisiae along a mixed-culture wine fermentation with Hanseniaspora guilliermondii
Abstract
Background: The introduction of yeast starter cultures consisting in a blend of Saccharomyces cerevisiae and non-Saccharomyces yeast strains is emerging for production of wines with improved complexity of flavor. The rational use of this approach is, however, dependent on knowing the impact that co-inoculation has in the physiology of S. cerevisiae. In this work the transcriptome of S. cerevisiae was monitored throughout a wine fermentation, carried out in single culture or in a consortium with Hanseniaspora guilliermondii, this being the first time that this relevant yeast-yeast interaction is examined at a genomic scale.
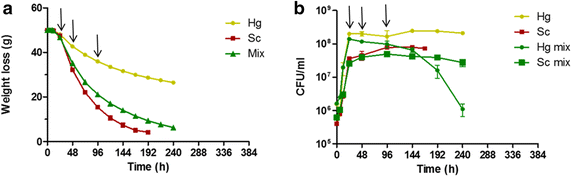
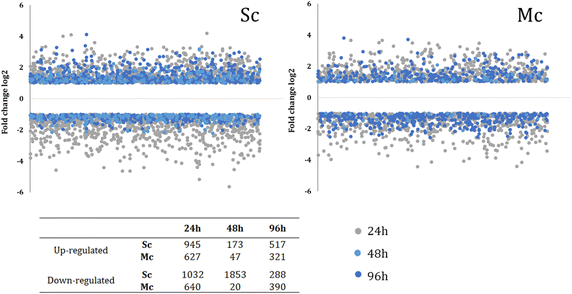
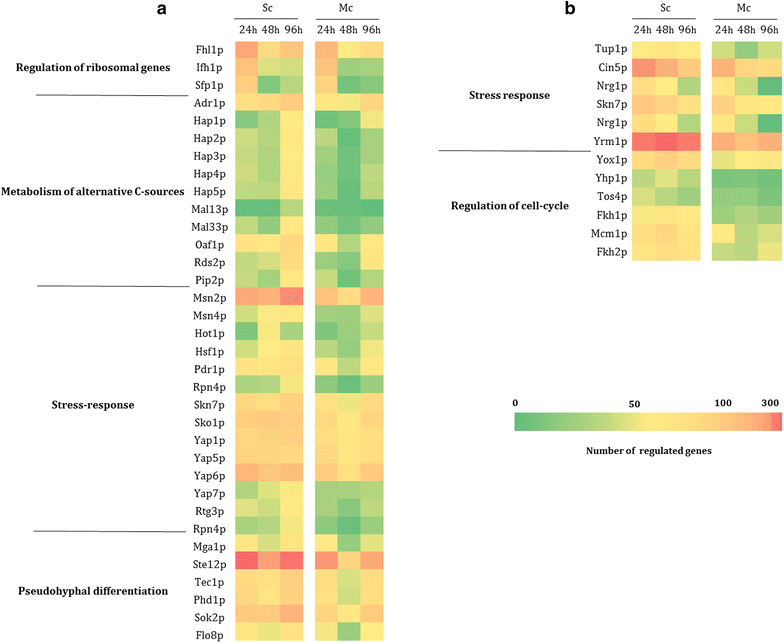
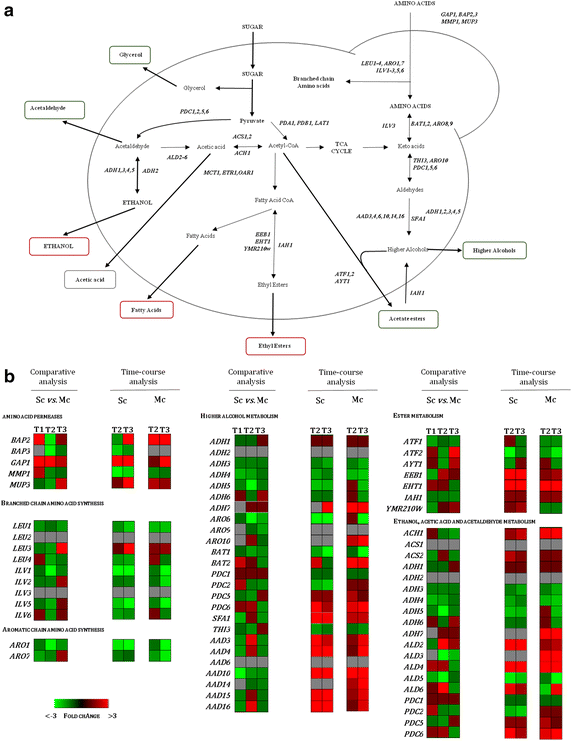
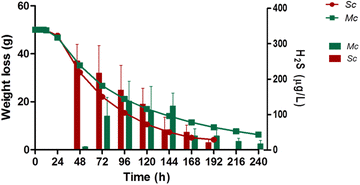
Results: Co-inoculation with H. guilliermondii reduced the overall genome-wide transcriptional response of S. cerevisiae throughout the fermentation, which was attributable to a lower fermentative activity of S. cerevisiae while in the mixed-fermentation. Approximately 350 genes S. cerevisiae genes were found to be differently expressed (FDR < 0.05) in response to the presence of H. guilliermondii in the fermentation medium. Genes involved in biosynthesis of vitamins were enriched among those up-regulated in the mixed-culture fermentation, while genes related with the uptake and biosynthesis of amino acids were enriched among those more expressed in the single-culture. The differences in the aromatic profiles of wines obtained in the single and in the mixed-fermentations correlated with the differential expression of S. cerevisiae genes encoding enzymes required for formation of aroma compounds.
Conclusions: By integrating results obtained in the transcriptomic analysis performed with physiological data our study provided, for the first time, an integrated view into the adaptive responses of S. cerevisiae to the challenging environment of mixed culture fermentation. The availability of nutrients, in particular, of nitrogen and vitamins, stands out as a factor that may determine population dynamics, fermentative activity and by-product formation.
Figures

References
-
- Jolly N, Augustyn O, Pretorius IS. The role and use of non-Saccharomyces yeasts in wine production. S Afr J Enol Vitic. 2006;27:15–39.
-
- Mora J, Barbas JI, Mulet A. Growth of yeast species during the fermentation of musts inoculated with Kluyveromyces thermotolerans and Saccharomycescerevisiae. Am J Enol Vitic. 1990;39:156–159.
-
- Soden A, Francis IL, Oakey H, Henschke PA. Effects of co-fermentation with Candida stellata and S. cerevisiae on the aroma and composition of Chardonnay wine. Aust J Grape Wine Res. 2000;6:21–30. doi: 10.1111/j.1755-0238.2000.tb00158.x. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

