Temporal variation selects for diet-microbe co-metabolic traits in the gut of Gorilla spp
- PMID: 26315972
- PMCID: PMC4737941
- DOI: 10.1038/ismej.2015.146
Temporal variation selects for diet-microbe co-metabolic traits in the gut of Gorilla spp
Erratum in
-
Temporal variation selects for diet-microbe co-metabolic traits in the gut of Gorilla spp.ISME J. 2016 Feb;10(2):532. doi: 10.1038/ismej.2015.252. ISME J. 2016. PMID: 26784513 Free PMC article. No abstract available.
Abstract
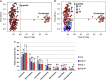
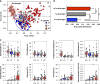
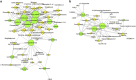
Although the critical role that our gastrointestinal microbes play in host physiology is now well established, we know little about the factors that influenced the evolution of primate gut microbiomes. To further understand current gut microbiome configurations and diet-microbe co-metabolic fingerprints in primates, from an evolutionary perspective, we characterized fecal bacterial communities and metabolomic profiles in 228 fecal samples of lowland and mountain gorillas (G. g. gorilla and G. b. beringei, respectively), our closest evolutionary relatives after chimpanzees. Our results demonstrate that the gut microbiomes and metabolomes of these two species exhibit significantly different patterns. This is supported by increased abundance of metabolites and bacterial taxa associated with fiber metabolism in mountain gorillas, and enrichment of markers associated with simple sugar, lipid and sterol turnover in the lowland species. However, longitudinal sampling shows that both species' microbiomes and metabolomes converge when hosts face similar dietary constraints, associated with low fruit availability in their habitats. By showing differences and convergence of diet-microbe co-metabolic fingerprints in two geographically isolated primate species, under specific dietary stimuli, we suggest that dietary constraints triggered during their adaptive radiation were potential factors behind the species-specific microbiome patterns observed in primates today.
Figures

References
-
- Amato KR, Leigh SR, Kent A, Mackie RI, Yeoman CJ, Stumpf RM et al. (2014. a). The gut microbiota appears to compensate for seasonal diet variation in the wild Black Howler Monkey (Alouatta pigra). Microb Ecol 69: 434–443. - PubMed
-
- Amato KR, Leigh SR, Kent A, Mackie RI, Yeoman CJ, Stumpf RM et al. (2014. b). The role of gut microbes in satisfying the nutritional demands of adult and juvenile wild, black howler monkeys (Alouatta pigra). Am J Phys Anthropol 155: 652–664. - PubMed
-
- Behrensmeyer AK. (2006). Climate change and human evolution. Science(Washington) 311: 476–478. - PubMed
-
- Chapman C, Rothman J, Lambert J. (2012) Food as a selective force in primates. The evolution of primate societies. University Of Chicago Press: Chicago, pp 149–168.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

