Modeling and Classification of Kinetic Patterns of Dynamic Metabolic Biomarkers in Physical Activity
- PMID: 26317529
- PMCID: PMC4552566
- DOI: 10.1371/journal.pcbi.1004454
Modeling and Classification of Kinetic Patterns of Dynamic Metabolic Biomarkers in Physical Activity
Abstract
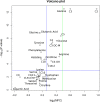
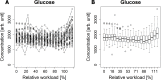
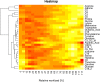
The objectives of this work were the classification of dynamic metabolic biomarker candidates and the modeling and characterization of kinetic regulatory mechanisms in human metabolism with response to external perturbations by physical activity. Longitudinal metabolic concentration data of 47 individuals from 4 different groups were examined, obtained from a cycle ergometry cohort study. In total, 110 metabolites (within the classes of acylcarnitines, amino acids, and sugars) were measured through a targeted metabolomics approach, combining tandem mass spectrometry (MS/MS) with the concept of stable isotope dilution (SID) for metabolite quantitation. Biomarker candidates were selected by combined analysis of maximum fold changes (MFCs) in concentrations and P-values resulting from statistical hypothesis testing. Characteristic kinetic signatures were identified through a mathematical modeling approach utilizing polynomial fitting. Modeled kinetic signatures were analyzed for groups with similar behavior by applying hierarchical cluster analysis. Kinetic shape templates were characterized, defining different forms of basic kinetic response patterns, such as sustained, early, late, and other forms, that can be used for metabolite classification. Acetylcarnitine (C2), showing a late response pattern and having the highest values in MFC and statistical significance, was classified as late marker and ranked as strong predictor (MFC = 1.97, P < 0.001). In the class of amino acids, highest values were shown for alanine (MFC = 1.42, P < 0.001), classified as late marker and strong predictor. Glucose yields a delayed response pattern, similar to a hockey stick function, being classified as delayed marker and ranked as moderate predictor (MFC = 1.32, P < 0.001). These findings coincide with existing knowledge on central metabolic pathways affected in exercise physiology, such as β-oxidation of fatty acids, glycolysis, and glycogenolysis. The presented modeling approach demonstrates high potential for dynamic biomarker identification and the investigation of kinetic mechanisms in disease or pharmacodynamics studies using MS data from longitudinal cohort studies.
Conflict of interest statement
KMW holds stock in sAnalytiCo Ltd. and in Biocrates Life Sciences AG. CB, MB, MN declare no competing interests.
Figures




References
-
- Guldberg CM, Waage P. Studier i affiniteten. (Translation: Studies on affinities.) Forhandlinger i Videnskabs-Selskabet i Christiania; 1864.
-
- Guldberg CM, Waage P. E´tudes sur les affinites chimiques. (Translation: Studies on chemical affinities.) Christiania: Brøgger & Christie; 1867.
-
- Guldberg CM, Waage P. Über die chemische Affinität. (Translation: On chemical affinity.) Erdmann’s Journal für practische Cehmie. 1879;127:69–114.
-
- Michaelis L, Menten ML. Die Kinetik der Invertinwirkung. (Translation: The kinetics of invertase activity.) Biochemische Zeitschrift. 1913;49:333–369.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

