Structural Basis for Substrate Specificity in Adenosylcobalamin-dependent Isobutyryl-CoA Mutase and Related Acyl-CoA Mutases
- PMID: 26318610
- PMCID: PMC4646380
- DOI: 10.1074/jbc.M115.676890
Structural Basis for Substrate Specificity in Adenosylcobalamin-dependent Isobutyryl-CoA Mutase and Related Acyl-CoA Mutases
Abstract
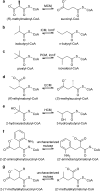
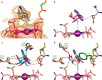
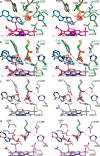
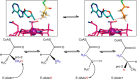
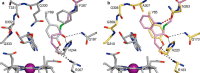
Acyl-CoA mutases are a growing class of adenosylcobalamin-dependent radical enzymes that perform challenging carbon skeleton rearrangements in primary and secondary metabolism. Members of this class of enzymes must precisely control substrate positioning to prevent oxidative interception of radical intermediates during catalysis. Our understanding of substrate specificity and catalysis in acyl-CoA mutases, however, is incomplete. Here, we present crystal structures of IcmF, a natural fusion protein variant of isobutyryl-CoA mutase, in complex with the adenosylcobalamin cofactor and four different acyl-CoA substrates. These structures demonstrate how the active site is designed to accommodate the aliphatic acyl chains of each substrate. The structures suggest that a conformational change of the 5'-deoxyadenosyl group from C2'-endo to C3'-endo could contribute to initiation of catalysis. Furthermore, detailed bioinformatic analyses guided by our structural findings identify critical determinants of acyl-CoA mutase substrate specificity and predict new acyl-CoA mutase-catalyzed reactions. These results expand our understanding of the substrate specificity and the catalytic scope of acyl-CoA mutases and could benefit engineering efforts for biotechnological applications ranging from production of biofuels and commercial products to hydrocarbon remediation.
Keywords: acyl-CoA mutase; adenosylcobalamin (AdoCbl); enzyme catalysis; metalloenzyme; substrate specificity; x-ray crystallography.
© 2015 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures

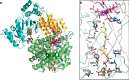
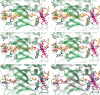




References
-
- Halpern J. (1985) Mechanisms of coenzyme B12-dependent rearrangements. Science 227, 869–875 - PubMed
-
- Frey P. A. (2010) in Comprehensive Natural Products II Chemistry and Biology (Mander L., and Liu H. W., eds) pp. 501–546, Elsevier, Oxford, UK
-
- Banerjee R. (2003) Radical carbon skeleton rearrangements: catalysis by coenzyme B12-dependent mutases. Chem. Rev. 103, 2083–2094 - PubMed
-
- Cracan V., and Banerjee R. (2013) Cobalt and corrinoid transport and biochemistry. Met. Ions Life Sci. 12, 333–374 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

