Metabolism of Fructooligosaccharides in Lactobacillus plantarum ST-III via Differential Gene Transcription and Alteration of Cell Membrane Fluidity
- PMID: 26319882
- PMCID: PMC4616932
- DOI: 10.1128/AEM.02426-15
Metabolism of Fructooligosaccharides in Lactobacillus plantarum ST-III via Differential Gene Transcription and Alteration of Cell Membrane Fluidity
Abstract
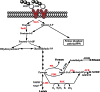
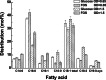
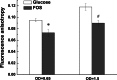
Although fructooligosaccharides (FOS) can selectively stimulate the growth and activity of probiotics and beneficially modulate the balance of intestinal microbiota, knowledge of the molecular mechanism for FOS metabolism by probiotics is still limited. Here a combined transcriptomic and physiological approach was used to survey the global alterations that occurred during the logarithmic growth of Lactobacillus plantarum ST-III using FOS or glucose as the sole carbon source. A total of 363 genes were differentially transcribed; in particular, two gene clusters were induced by FOS. Gene inactivation revealed that both of the clusters participated in the metabolism of FOS, which were transported across the membrane by two phosphotransferase systems (PTSs) and were subsequently hydrolyzed by a β-fructofuranosidase (SacA) in the cytoplasm. Combining the measurements of the transcriptome- and membrane-related features, we discovered that the genes involved in the biosynthesis of fatty acids (FAs) were repressed in cells grown on FOS; as a result, the FA profiles were altered by shortening of the carbon chains, after which membrane fluidity increased in response to FOS transport and utilization. Furthermore, incremental production of acetate was observed in both the transcriptomic and the metabolic experiments. Our results provided new insights into gene transcription, the production of metabolites, and membrane alterations that could explain FOS metabolism in L. plantarum.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures




References
-
- Gibson GR, Roberfroid MB. 1995. Dietary modulation of the human colonic microbiota: introducing the concept of prebiotics. J Nutr 125:1401–1412. - PubMed
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

