Spatial regulation of cytoplasmic snRNP assembly at the cellular level
- PMID: 26320237
- PMCID: PMC4765780
- DOI: 10.1093/jxb/erv399
Spatial regulation of cytoplasmic snRNP assembly at the cellular level
Abstract
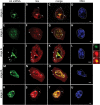
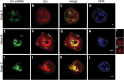
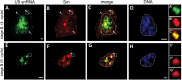
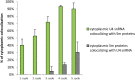
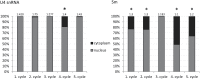
Small nuclear ribonucleoproteins (snRNPs) play a crucial role in pre-mRNA splicing in all eukaryotic cells. In contrast to the relatively broad knowledge on snRNP assembly within the nucleus, the spatial organization of the cytoplasmic stages of their maturation remains poorly understood. Nevertheless, sparse research indicates that, similar to the nuclear steps, the crucial processes of cytoplasmic snRNP assembly may also be strictly spatially regulated. In European larch microsporocytes, it was determined that the cytoplasmic assembly of snRNPs within a cell might occur in two distinct spatial manners, which depend on the rate of de novo snRNP formation in relation to the steady state of these particles within the nucleus. During periods of moderate expression of splicing elements, the cytoplasmic assembly of snRNPs occurred diffusely throughout the cytoplasm. Increased expression of both Sm proteins and U snRNA triggered the accumulation of these particles within distinct, non-membranous RNP-rich granules, which are referred to as snRNP-rich cytoplasmic bodies.
Keywords: Cajal bodies; Larix; P bodies; Sm proteins; confocal microscopy; in situ hybridization; plant cell; snRNA; splicing..
© The Author 2015. Published by Oxford University Press on behalf of the Society for Experimental Biology.
Figures

Similar articles
-
Periodic expression of Sm proteins parallels formation of nuclear Cajal bodies and cytoplasmic snRNP-rich bodies.Histochem Cell Biol. 2011 Nov;136(5):527-41. doi: 10.1007/s00418-011-0861-8. Epub 2011 Sep 9. Histochem Cell Biol. 2011. PMID: 21904826 Free PMC article.
-
A role for Cajal bodies in the final steps of U2 snRNP biogenesis.J Cell Sci. 2004 Sep 1;117(Pt 19):4423-33. doi: 10.1242/jcs.01308. Epub 2004 Aug 17. J Cell Sci. 2004. PMID: 15316075
-
U bodies are cytoplasmic structures that contain uridine-rich small nuclear ribonucleoproteins and associate with P bodies.Proc Natl Acad Sci U S A. 2007 Jul 10;104(28):11655-9. doi: 10.1073/pnas.0704977104. Epub 2007 Jun 26. Proc Natl Acad Sci U S A. 2007. PMID: 17595295 Free PMC article.
-
Plant snRNP Biogenesis: A Perspective from the Nucleolus and Cajal Bodies.Front Plant Sci. 2018 Jan 4;8:2184. doi: 10.3389/fpls.2017.02184. eCollection 2017. Front Plant Sci. 2018. PMID: 29354141 Free PMC article. Review.
-
Assembly and intracellular transport of snRNP particles.Bioessays. 1991 Feb;13(2):57-64. doi: 10.1002/bies.950130203. Bioessays. 1991. PMID: 1827581 Review.
Cited by
-
Spatio-temporal localization of LlBOP following early events of floral abscission in yellow lupine.Protoplasma. 2019 Sep;256(5):1173-1183. doi: 10.1007/s00709-019-01365-3. Epub 2019 Apr 16. Protoplasma. 2019. PMID: 30993471 Free PMC article.
-
Core spliceosomal Sm proteins as constituents of cytoplasmic mRNPs in plants.Plant J. 2020 Aug;103(3):1155-1173. doi: 10.1111/tpj.14792. Epub 2020 May 28. Plant J. 2020. PMID: 32369637 Free PMC article.
-
Cajal bodies: Evolutionarily conserved nuclear biomolecular condensates with properties unique to plants.Plant Cell. 2023 Sep 1;35(9):3214-3235. doi: 10.1093/plcell/koad140. Plant Cell. 2023. PMID: 37202374 Free PMC article. Review.
-
Non-ubiquitous expression of core spliceosomal protein SmB/B' in chick and mouse embryos.Dev Dyn. 2023 Feb;252(2):276-293. doi: 10.1002/dvdy.537. Epub 2022 Sep 20. Dev Dyn. 2023. PMID: 36058892 Free PMC article.
-
Functional nuclear retention of pre-mRNA involving Cajal bodies during meiotic prophase in European larch (Larix decidua).Plant Cell. 2022 May 24;34(6):2404-2423. doi: 10.1093/plcell/koac091. Plant Cell. 2022. PMID: 35294035 Free PMC article.
References
-
- Barbee SA, Lublin AL, Evans TC. 2002. A novel function for the Sm proteins in germ granule localization during C. elegans embryogenesis. Current Biology 12, 1502–1506. - PubMed
-
- Baserga SJ, Steitz JA. 1993. The diverse world of small ribonucleoproteins. In: Gesteland RF, Atkins JF, eds. The RNA World. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press, 359–381.
-
- Bednarska E. 1984. Ultrastructural and metabolic transformations of differentiating Hyacinthus orientalis L., pollen grain cells. I. RNA and protein synthesis. Acta Societatis Botanicorum Poloniae 53, 145–158.
-
- Bilinski SM, Jaglarz MK, Szymanska B, Etkin LD, Kloc M. 2004. Sm proteins, the constituents of the spliceosome, are components of nuage and mitochondrial cement in Xenopus oocytes. Experimental Cell Research 299, 171–178. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

