TNF-α-Induced microRNAs Control Dystrophin Expression in Becker Muscular Dystrophy
- PMID: 26321630
- PMCID: PMC4757433
- DOI: 10.1016/j.celrep.2015.07.066
TNF-α-Induced microRNAs Control Dystrophin Expression in Becker Muscular Dystrophy
Abstract
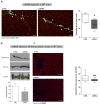
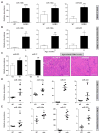
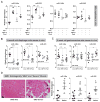
The amount and distribution of dystrophin protein in myofibers and muscle is highly variable in Becker muscular dystrophy and in exon-skipping trials for Duchenne muscular dystrophy. Here, we investigate a molecular basis for this variability. In muscle from Becker patients sharing the same exon 45-47 in-frame deletion, dystrophin levels negatively correlate with microRNAs predicted to target dystrophin. Seven microRNAs inhibit dystrophin expression in vitro, and three are validated in vivo (miR-146b/miR-374a/miR-31). microRNAs are expressed in dystrophic myofibers and increase with age and disease severity. In exon-skipping-treated mdx mice, microRNAs are significantly higher in muscles with low dystrophin rescue. TNF-α increases microRNA levels in vitro whereas NFκB inhibition blocks this in vitro and in vivo. Collectively, these data show that microRNAs contribute to variable dystrophin levels in muscular dystrophy. Our findings suggest a model where chronic inflammation in distinct microenvironments induces pathological microRNAs, initiating a self-sustaining feedback loop that exacerbates disease progression.
Copyright © 2015 The Authors. Published by Elsevier Inc. All rights reserved.
Figures



References
-
- Calabia-Linares C, Robles-Valero J, de la Fuente H, Perez-Martinez M, Martin-Cofreces N, Alfonso-Perez M, Gutierrez-Vazquez C, Mittelbrunn M, Ibiza S, Urbano-Olmos FR, et al. Endosomal clathrin drives actin accumulation at the immunological synapse. J Cell Sci. 2011;124:820–830. - PubMed
-
- Chamberlain JS, Grant SG, Reeves AA, Mullins LJ, Stephenson DA, Hoffman EP, Monaco AP, Kunkel LM, Caskey CT, Chapman VM. Regional localization of the murine Duchenne muscular dystrophy gene on the mouse X chromosome. Somat Cell Mol Genet. 1987;13:671–678. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- T32 AR056993/AR/NIAMS NIH HHS/United States
- L40 AR068727/AR/NIAMS NIH HHS/United States
- 5T32AR056993/AR/NIAMS NIH HHS/United States
- GTB12001/TI_/Telethon/Italy
- K99 HL130035/HL/NHLBI NIH HHS/United States
- R00 HL130035/HL/NHLBI NIH HHS/United States
- R01 NS029525/NS/NINDS NIH HHS/United States
- 1P50AR060836-01/AR/NIAMS NIH HHS/United States
- 1U54HD071601-01/HD/NICHD NIH HHS/United States
- K99HL130035/HL/NHLBI NIH HHS/United States
- P50 AR060836/AR/NIAMS NIH HHS/United States
- U54 HD071601/HD/NICHD NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

