Profiling of promoter occupancy by the SND1 transcriptional coactivator identifies downstream glycerolipid metabolic genes involved in TNFα response in human hepatoma cells
- PMID: 26323317
- PMCID: PMC4678849
- DOI: 10.1093/nar/gkv858
Profiling of promoter occupancy by the SND1 transcriptional coactivator identifies downstream glycerolipid metabolic genes involved in TNFα response in human hepatoma cells
Abstract
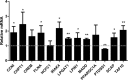
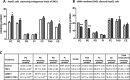
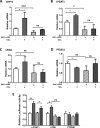
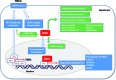
The NF-κB-inducible Staphylococcal nuclease and tudor domain-containing 1 gene (SND1) encodes a coactivator involved in inflammatory responses and tumorigenesis. While SND1 is known to interact with certain transcription factors and activate client gene expression, no comprehensive mapping of SND1 target genes has been reported. Here, we have approached this question by performing ChIP-chip assays on human hepatoma HepG2 cells and analyzing SND1 binding modulation by proinflammatory TNFα. We show that SND1 binds 645 gene promoters in control cells and 281 additional genes in TNFα-treated cells. Transcription factor binding site analysis of bound probes identified motifs for established partners and for novel transcription factors including HSF, ATF, STAT3, MEIS1/AHOXA9, E2F and p300/CREB. Major target genes were involved in gene expression and RNA metabolism regulation, as well as development and cellular metabolism. We confirmed SND1 binding to 21 previously unrecognized genes, including a set of glycerolipid genes. Knocking-down experiments revealed that SND1 deficiency compromises the glycerolipid gene reprogramming and lipid phenotypic responses to TNFα. Overall, our findings uncover an unexpected large set of potential SND1 target genes and partners and reveal SND1 to be a determinant downstream effector of TNFα that contributes to support glycerophospholipid homeostasis in human hepatocellular carcinoma during inflammation.
© The Author(s) 2015. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


References
-
- Broadhurst M.K., Wheeler T.T. The p100 coactivator is present in the nuclei of mammary epithelial cells and its abundance is increased in response to prolactin in culture and in mammary tissue during lactation. J. Endocrinol. 2001;171:329–337. - PubMed
-
- Zhao C.T., Shi K.H., Su Y., Liang L.Y., Yan Y., Postlethwait J., Meng A.M. Two variants of zebrafish p100 are expressed during embryogenesis and regulated by nodal signaling. FEBS Lett. 2003;543:190–195. - PubMed
-
- Hossain M.J., Korde R., Singh P.K., Kanodia S., Ranjan R., Ram G., Kalsey G.S., Singh R., Malhotra P. Plasmodium falciparum tudor staphylococcal nuclease interacting proteins suggest its role in nuclear as well as splicing processes. Gene. 2010;468:48–57. - PubMed
-
- Rodriguez L., Ochoa B., Martinez M.J. NF-Y and Sp1 are involved in transcriptional regulation of rat SND p102 gene. Biochem. Biophys. Res. Commun. 2007;356:226–232. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

