Rooting the tree of life: the phylogenetic jury is still out
- PMID: 26323760
- PMCID: PMC4571568
- DOI: 10.1098/rstb.2014.0329
Rooting the tree of life: the phylogenetic jury is still out
Abstract
This article aims to shed light on difficulties in rooting the tree of life (ToL) and to explore the (sociological) reasons underlying the limited interest in accurately addressing this fundamental issue. First, we briefly review the difficulties plaguing phylogenetic inference and the ways to improve the modelling of the substitution process, which is highly heterogeneous, both across sites and over time. We further observe that enriched taxon samplings, better gene samplings and clever data removal strategies have led to numerous revisions of the ToL, and that these improved shallow phylogenies nearly always relocate simple organisms higher in the ToL provided that long-branch attraction artefacts are kept at bay. Then, we note that, despite the flood of genomic data available since 2000, there has been a surprisingly low interest in inferring the root of the ToL. Furthermore, the rare studies dealing with this question were almost always based on methods dating from the 1990s that have been shown to be inaccurate for much more shallow issues! This leads us to argue that the current consensus about a bacterial root for the ToL can be traced back to the prejudice of Aristotle's Great Chain of Beings, in which simple organisms are ancestors of more complex life forms. Finally, we demonstrate that even the best models cannot yet handle the complexity of the evolutionary process encountered both at shallow depth, when the outgroup is too distant, and at the level of the inter-domain relationships. Altogether, we conclude that the commonly accepted bacterial root is still unproven and that the root of the ToL should be revisited using phylogenomic supermatrices to ensure that new evidence for eukaryogenesis, such as the recently described Lokiarcheota, is interpreted in a sound phylogenetic framework.
Keywords: Great Chain of Beings prejudice; model of evolution; simplification; systematic error; tree reconstruction artefact.
© 2015 The Author(s).
Figures

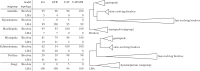

References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
