Effective Small Interfering RNA Therapy to Treat CLCN7-dependent Autosomal Dominant Osteopetrosis Type 2
- PMID: 26325626
- PMCID: PMC4877447
- DOI: 10.1038/mtna.2015.21
Effective Small Interfering RNA Therapy to Treat CLCN7-dependent Autosomal Dominant Osteopetrosis Type 2
Abstract
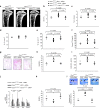
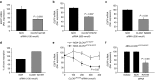
In about 70% of patients affected by autosomal dominant osteopetrosis type 2 (ADO2), osteoclast activity is reduced by heterozygous mutations of the CLCN7 gene, encoding the ClC-7 chloride/hydrogen antiporter. CLCN7(G215R)-, CLCN7(R767W)-, and CLCN7(R286W)-specific siRNAs silenced transfected mutant mRNA/EGFP in HEK293 cells, in RAW264.7 cells and in human osteoclasts, with no change of CLCN7(WT) mRNA and no effect of scrambled siRNA on the mutant transcripts. Osteoclasts from Clcn7(G213R) ADO2 mice showed reduced bone resorption, a condition rescued by Clcn7(G213R)-specific siRNA. Treatment of ADO2 mice with Clcn7(G213R)-specific siRNA induced increase of bone resorption variables and decrease of trabecular bone mass, leading to an overall improvement of the osteopetrotic bone phenotype. Treatment did not induce overt adverse effects and was effective also with siRNAs specific for other mutants. These results demonstrate that a siRNA-based experimental treatment of ADO2 is feasible, and underscore a translational impact for future strategy to cure this therapeutically neglected form of osteopetrosis.
Figures



References
-
- Anasagasti, A, Irigoyen, C, Barandika, O, López de Munain, A and Ruiz-Ederra, J (2012). Current mutation discovery approaches in Retinitis Pigmentosa. Vision Res 75: 117–129. - PubMed
-
- Del Fattore, A, Cappariello, A and Teti, A (2008). Genetics, pathogenesis and complications of osteopetrosis. Bone 42: 19–29. - PubMed
-
- Lu, X, Yang, X, Huang, X, Huang, C, Sun, HH, Jin, L et al. (2013). RNA interference targeting E637K mutation rescues hERG channel currents and restores its kinetic properties. Heart Rhythm 10: 128–136. - PubMed
-
- Leslie Pedrioli, DM, Fu, DJ, Gonzalez-Gonzalez, E, Contag, CH, Kaspar, RL, Smith, FJ et al. (2012). Generic and personalized RNAi-based therapeutics for a dominant-negative epidermal fragility disorder. J Invest Dermatol 132: 1627–1635. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

