Elaborate uORF/IRES features control expression and localization of human glycyl-tRNA synthetase
- PMID: 26327585
- PMCID: PMC4829322
- DOI: 10.1080/15476286.2015.1086866
Elaborate uORF/IRES features control expression and localization of human glycyl-tRNA synthetase
Abstract
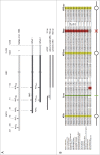
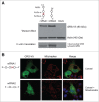
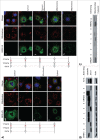
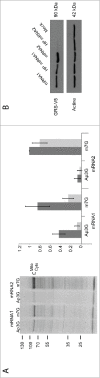
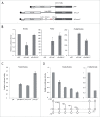
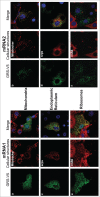
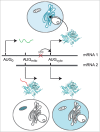
The canonical activity of glycyl-tRNA synthetase (GARS) is to charge glycine onto its cognate tRNAs. However, outside translation, GARS also participates in many other functions. A single gene encodes both the cytosolic and mitochondrial forms of GARS but 2 mRNA isoforms were identified. Using immunolocalization assays, in vitro translation assays and bicistronic constructs we provide experimental evidence that one of these mRNAs tightly controls expression and localization of human GARS. An intricate regulatory domain was found in its 5'-UTR which displays a functional Internal Ribosome Entry Site and an upstream Open Reading Frame. Together, these elements hinder the synthesis of the mitochondrial GARS and target the translation of the cytosolic enzyme to ER-bound ribosomes. This finding reveals a complex picture of GARS translation and localization in mammals. In this context, we discuss how human GARS expression could influence its moonlighting activities and its involvement in diseases.
Keywords: IRES; aminoacyl-tRNA synthetase; post-transcriptional control; uORF.
Figures
References
-
- Park SG, Ewalt KL, Kim S. Functional expansion of aminoacyl-tRNA synthetases and their interacting factors: new perspectives on housekeepers. Trends Biochem Sci 2005; 30:569-74; PMID:16125937; http://dx.doi.org/ 10.1016/j.tibs.2005.08.004 - DOI - PubMed
-
- Hausmann CD, Ibba M. Aminoacyl-tRNA synthetase complexes: molecular multitasking revealed. FEMS Microbiol Rev 2008; 32:705-21; PMID:18522650; http://dx.doi.org/ 10.1111/j.1574-6976.2008.00119.x - DOI - PMC - PubMed
-
- Kim S, You S, Hwang D. Aminoacyl-tRNA synthetases and tumorigenesis: more than housekeeping. Nat Rev Cancer 2011; 11:708-18; PMID:21941282; http://dx.doi.org/ 10.1038/nrc3124 - DOI - PubMed
-
- Guo M, Schimmel P. Essential nontranslational functions of tRNA synthetases. Nat Chem Biol 2013; 9:145-53; PMID:23416400; http://dx.doi.org/ 10.1038/nchembio.1158 - DOI - PMC - PubMed
-
- Antonellis A, Ellsworth RE, Sambuughin N, Puls I, Abel A, Lee-Lin SQ, Jordanova A, Kremensky I, Christodoulou K, Middleton LT, et al.. Glycyl tRNA synthetase mutations in Charcot-Marie-Tooth disease type 2D and distal spinal muscular atrophy type V. Am J Hum Genet 2003; 72:1293-9; PMID:12690580; http://dx.doi.org/ 10.1086/375039 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
