Translational metabolomics in cancer research
- PMID: 26329905
- PMCID: PMC4718447
- DOI: 10.2217/bmm.15.52
Translational metabolomics in cancer research
Abstract
Over the last decade there has been a bottleneck in the introduction of new validated cancer metabolic biomarkers into clinical practice. Unfortunately, there are no biomarkers with adequate sensitivity for the early detection of cancer, and there remain a reliance on cancer antigens for monitoring treatment. The need for new diagnostics has led to the exploration of untargeted metabolomics for discovery of early biomarkers of specific cancers and targeted metabolomics to elucidate mechanistic aspects of tumor progression. The successful translation of such strategies to the treatment of cancer would allow earlier intervention to improve survival. We have reviewed the methodology that is being used to achieve these goals together with recent advances in implementing translational metabolomics in cancer.
Keywords: NMR spectroscopy; cancer; cancer diagnosis; diagnostic biomarkers; lipidomics; liquid chromatography-mass spectrometry; prognostic biomarkers; untargeted metabolomics targeted metabolomics.
Conflict of interest statement
The authors have no other relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
No writing assistance was utilized in the production of this manuscript.
Figures
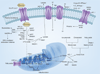
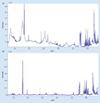
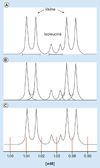



References
-
- O’Connell TM. Recent advances in metabolomics in oncology. Bioanalysis. 2012;4(4):431–451. - PubMed
-
- Blair IA. Eliminating Bottlenecks for Efficient Bioanalysis: Practices and Applications in Drug Discovery and Development. London, UK: Future Science Group; 2015. Bioanalysis supporting disease biomarker discovery and validation; pp. 162–181.
-
- Forcisi S, Moritz F, Kanawati B, Tziotis D, Lehmann R, Schmitt-Kopplin P. Liquid chromatography-mass spectrometry in metabolomics research: mass analyzers in ultra high pressure liquid chromatography coupling. J. Chromatogr. A. 2013;1292:51–65. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
