A quantitative proteomics analysis of MCF7 breast cancer stem and progenitor cell populations
- PMID: 26332018
- PMCID: PMC4715579
- DOI: 10.1002/pmic.201500002
A quantitative proteomics analysis of MCF7 breast cancer stem and progenitor cell populations
Abstract
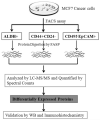
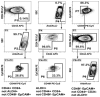
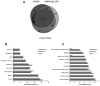
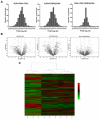
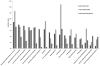
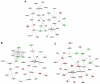
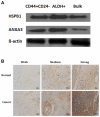
Accumulating evidence has demonstrated that breast cancers are initiated and develop from a small population of stem-like cells termed cancer stem cells (CSCs). These cells are hypothesized to mediate tumor metastasis and contribute to therapeutic resistance. However, the molecular regulatory networks responsible for maintaining CSCs in an undifferentiated state have yet to be elucidated. In this study, we used CSC markers to isolate pure breast CSCs fractions (ALDH+ and CD44+CD24- cell populations) and the mature luminal cells (CD49f-EpCAM+) from the MCF7 cell line. Proteomic analysis was performed on these samples and a total of 3304 proteins were identified. A label-free quantitative method was applied to analyze differentially expressed proteins. Using the criteria of greater than twofold changes and p value <0.05, 305, 322 and 98 proteins were identified as significantly different in three pairwise comparisons of ALDH+ versus CD44+CD24-, ALDH+ versus CD49f-EpCAM+ and CD44+CD24- versus CD49f-EpCAM+, respectively. Pathway analysis of differentially expressed proteins by Ingenuity Pathway Analysis (IPA) revealed potential molecular regulatory networks that may regulate CSCs. Selected differential proteins were validated by Western blot assay and immunohistochemical staining. The use of proteomics analysis may increase our understanding of the underlying molecular mechanisms of breast CSCs. This may be of importance in the future development of anti-CSC therapeutics.
Keywords: Breast cancer; Cancer stem cell; LC-MS/MS; Pathway analysis; Quantitative proteomics; Systems biology.
© 2015 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Figures
Similar articles
-
Single Amino Acid Variant Profiles of Subpopulations in the MCF-7 Breast Cancer Cell Line.J Proteome Res. 2017 Feb 3;16(2):842-851. doi: 10.1021/acs.jproteome.6b00824. Epub 2017 Jan 20. J Proteome Res. 2017. PMID: 28076950 Free PMC article.
-
Semi-quantitative evaluation of CD44(+) /CD24(-) tumor cell distribution in breast cancer tissue using a newly developed fluorescence immunohistochemical staining method.Cancer Sci. 2011 Dec;102(12):2132-8. doi: 10.1111/j.1349-7006.2011.02063.x. Epub 2011 Sep 14. Cancer Sci. 2011. PMID: 21838786
-
CD49f-positive cell population efficiently enriches colon cancer-initiating cells.Int J Oncol. 2013 Aug;43(2):425-30. doi: 10.3892/ijo.2013.1955. Epub 2013 May 24. Int J Oncol. 2013. PMID: 23708747
-
Surface markers of hepatocellular cancer stem cells and their clinical potential.Neoplasma. 2014;61(5):505-13. doi: 10.4149/neo_2014_061. Neoplasma. 2014. PMID: 24712843 Review.
-
Stem cells in human breast cancer.Histol Histopathol. 2010 Mar;25(3):371-85. doi: 10.14670/HH-25.371. Histol Histopathol. 2010. PMID: 20054808 Review.
Cited by
-
Tumacrophage: macrophages transformed into tumor stem-like cells by virulent genetic material from tumor cells.Oncotarget. 2017 Jul 18;8(47):82326-82343. doi: 10.18632/oncotarget.19320. eCollection 2017 Oct 10. Oncotarget. 2017. PMID: 29137267 Free PMC article.
-
Sulforaphane enhances the anticancer activity of taxanes against triple negative breast cancer by killing cancer stem cells.Cancer Lett. 2017 May 28;394:52-64. doi: 10.1016/j.canlet.2017.02.023. Epub 2017 Feb 27. Cancer Lett. 2017. PMID: 28254410 Free PMC article.
-
Anti-tumour effects of a dual cancer-specific oncolytic adenovirus on Breast Cancer Stem cells.J Cell Mol Med. 2021 Jan;25(2):666-676. doi: 10.1111/jcmm.16113. Epub 2020 Dec 11. J Cell Mol Med. 2021. PMID: 33305893 Free PMC article.
-
Differential proteomic comparison of breast cancer secretome using a quantitative paired analysis workflow.BMC Cancer. 2019 Apr 18;19(1):365. doi: 10.1186/s12885-019-5547-y. BMC Cancer. 2019. PMID: 30999875 Free PMC article.
-
Cancer stem cell metabolism.Breast Cancer Res. 2016 May 24;18(1):55. doi: 10.1186/s13058-016-0712-6. Breast Cancer Res. 2016. PMID: 27220421 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

