The structural and functional effects of the familial hypertrophic cardiomyopathy-linked cardiac troponin C mutation, L29Q
- PMID: 26341255
- PMCID: PMC4640586
- DOI: 10.1016/j.yjmcc.2015.08.017
The structural and functional effects of the familial hypertrophic cardiomyopathy-linked cardiac troponin C mutation, L29Q
Abstract
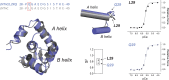
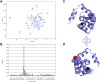
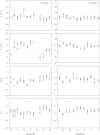
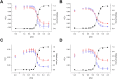
Familial hypertrophic cardiomyopathy (FHC) is characterized by severe abnormal cardiac muscle growth. The traditional view of disease progression in FHC is that an increase in the Ca(2+)-sensitivity of cardiac muscle contraction ultimately leads to pathogenic myocardial remodeling, though recent studies suggest this may be an oversimplification. For example, FHC may be developed through altered signaling that prevents downstream regulation of contraction. The mutation L29Q, found in the Ca(2+)-binding regulatory protein in heart muscle, cardiac troponin C (cTnC), has been linked to cardiac hypertrophy. However, reports on the functional effects of this mutation are conflicting, and our goal was to combine in vitro and in situ structural and functional data to elucidate its mechanism of action. We used nuclear magnetic resonance and circular dichroism to solve the structure and characterize the backbone dynamics and stability of the regulatory domain of cTnC with the L29Q mutation. The overall structure and dynamics of cTnC were unperturbed, although a slight rearrangement of site 1, an increase in backbone flexibility, and a small decrease in protein stability were observed. The structure and function of cTnC was also assessed in demembranated ventricular trabeculae using fluorescence for in situ structure. L29Q reduced the cooperativity of the Ca(2+)-dependent structural change in cTnC in trabeculae under basal conditions and abolished the effect of force-generating myosin cross-bridges on this structural change. These effects could contribute to the pathogenesis of this mutation.
Keywords: Cardiac; Fluorescence spectroscopy; Hypertrophic cardiomyopathy; L29Q; NMR spectroscopy; Troponin C.
Copyright © 2015 The Authors. Published by Elsevier Ltd.. All rights reserved.
Figures




Similar articles
-
Familial hypertrophic cardiomyopathy related cardiac troponin C L29Q mutation alters length-dependent activation and functional effects of phosphomimetic troponin I*.PLoS One. 2013 Nov 18;8(11):e79363. doi: 10.1371/journal.pone.0079363. eCollection 2013. PLoS One. 2013. PMID: 24260207 Free PMC article.
-
Structural kinetics of cardiac troponin C mutants linked to familial hypertrophic and dilated cardiomyopathy in troponin complexes.J Biol Chem. 2008 Feb 8;283(6):3424-3432. doi: 10.1074/jbc.M703822200. Epub 2007 Dec 5. J Biol Chem. 2008. PMID: 18063575
-
Familial hypertrophic cardiomyopathy-related cardiac troponin C mutation L29Q affects Ca2+ binding and myofilament contractility.Physiol Genomics. 2008 Apr 22;33(2):257-66. doi: 10.1152/physiolgenomics.00154.2007. Epub 2008 Feb 19. Physiol Genomics. 2008. PMID: 18285522
-
Structure and function of cardiac troponin C (TNNC1): Implications for heart failure, cardiomyopathies, and troponin modulating drugs.Gene. 2015 Oct 25;571(2):153-66. doi: 10.1016/j.gene.2015.07.074. Epub 2015 Jul 29. Gene. 2015. PMID: 26232335 Free PMC article. Review.
-
Biology of the troponin complex in cardiac myocytes.Prog Cardiovasc Dis. 2004 Nov-Dec;47(3):159-76. doi: 10.1016/j.pcad.2004.07.003. Prog Cardiovasc Dis. 2004. PMID: 15736582 Review.
Cited by
-
Cycling Cross-Bridges Contribute to Thin Filament Activation in Human Slow-Twitch Fibers.Front Physiol. 2020 Mar 24;11:144. doi: 10.3389/fphys.2020.00144. eCollection 2020. Front Physiol. 2020. PMID: 32265723 Free PMC article.
-
Calcium Handling in Inherited Cardiac Diseases: A Focus on Catecholaminergic Polymorphic Ventricular Tachycardia and Hypertrophic Cardiomyopathy.Int J Mol Sci. 2023 Feb 8;24(4):3365. doi: 10.3390/ijms24043365. Int J Mol Sci. 2023. PMID: 36834774 Free PMC article. Review.
-
Exploring contractile protein mechanisms and target medications for cardiomyopathic patients with diastolic dysfunction.Pediatr Discov. 2024 Mar 25;2(1):e60. doi: 10.1002/pdi3.60. eCollection 2024 Mar. Pediatr Discov. 2024. PMID: 40626249 Free PMC article. Review.
-
Troponin structure and function: a view of recent progress.J Muscle Res Cell Motil. 2020 Mar;41(1):71-89. doi: 10.1007/s10974-019-09513-1. Epub 2019 Apr 27. J Muscle Res Cell Motil. 2020. PMID: 31030382 Free PMC article. Review.
-
A green tea extract catechin EGCg: Therapeutic potential for pediatric cardiomyopathies.Pediatr Discov. 2023 Jun 9;1(1):e7. doi: 10.1002/pdi3.7. eCollection 2023 Jun. Pediatr Discov. 2023. PMID: 40625575 Free PMC article. Review.
References
-
- Gordon A.M., Homsher E., Regnier M. Regulation of contraction in striated muscle. Physiol. Rev. 2000;80:853–924. - PubMed
-
- Li M.X., Wang X., Sykes B.D. Structural based insights into the role of troponin in cardiac muscle pathophysiology. J. Muscle Res. Cell Motil. 2004;25:559–579. - PubMed
-
- Parmacek M.S., Solaro R.J. Biology of the troponin complex in cardiac myocytes. Prog. Cardiovasc. Dis. 2004;47:159–176. - PubMed
-
- Willott R.H., Gomes A.V., Chang A.N., Parvatiyar M.S., Pinto J.R., Potter J.D. Mutations in troponin that cause HCM, DCM AND RCM: what can we learn about thin filament function? J. Mol. Cell. Cardiol. 2010;48:882–892. - PubMed
-
- Ashrafian H., McKenna W.J., Watkins H. Disease pathways and novel therapeutic targets in hypertrophic cardiomyopathy. Circ. Res. 2011;109:86–96. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

