Class II HLA interactions modulate genetic risk for multiple sclerosis
- PMID: 26343388
- PMCID: PMC4874245
- DOI: 10.1038/ng.3395
Class II HLA interactions modulate genetic risk for multiple sclerosis
Abstract
Association studies have greatly refined the understanding of how variation within the human leukocyte antigen (HLA) genes influences risk of multiple sclerosis. However, the extent to which major effects are modulated by interactions is poorly characterized. We analyzed high-density SNP data on 17,465 cases and 30,385 controls from 11 cohorts of European ancestry, in combination with imputation of classical HLA alleles, to build a high-resolution map of HLA genetic risk and assess the evidence for interactions involving classical HLA alleles. Among new and previously identified class II risk alleles (HLA-DRB1*15:01, HLA-DRB1*13:03, HLA-DRB1*03:01, HLA-DRB1*08:01 and HLA-DQB1*03:02) and class I protective alleles (HLA-A*02:01, HLA-B*44:02, HLA-B*38:01 and HLA-B*55:01), we find evidence for two interactions involving pairs of class II alleles: HLA-DQA1*01:01-HLA-DRB1*15:01 and HLA-DQB1*03:01-HLA-DQB1*03:02. We find no evidence for interactions between classical HLA alleles and non-HLA risk-associated variants and estimate a minimal effect of polygenic epistasis in modulating major risk alleles.
Figures



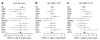
References
-
- Jersild C, Svejgaard A, Fog T. HL-A antigens and multiple sclerosis. Lancet. 1972;1:1240–1241. - PubMed
-
- Kwon OJ, et al. HLA class II susceptibility to multiple sclerosis among Ashkenazi and non-Ashkenazi Jews. Arch. Neurol. 1999;56:555–560. - PubMed
-
- International Multiple Sclerosis Genetics Consortium Risk alleles for multiple sclerosis identified by a genomewide study. N. Engl. J. Med. 2007;357:851–862. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01NS032830/NS/NINDS NIH HHS/United States
- R25 NS070680/NS/NINDS NIH HHS/United States
- 085475/B/08/Z/WT_/Wellcome Trust/United Kingdom
- R01 NS067305/NS/NINDS NIH HHS/United States
- 100956/WT_/Wellcome Trust/United Kingdom
- RC2GM093080/GM/NIGMS NIH HHS/United States
- RC2 GM093080/GM/NIGMS NIH HHS/United States
- RC2NS070340/NS/NINDS NIH HHS/United States
- 1UL1RR024975-01/RR/NCRR NIH HHS/United States
- NS049477/NS/NINDS NIH HHS/United States
- R01 NS026799/NS/NINDS NIH HHS/United States
- 098051/WT_/Wellcome Trust/United Kingdom
- 085475/Z/08/Z/WT_/Wellcome Trust/United Kingdom
- R01 NS049477/NS/NINDS NIH HHS/United States
- RC2 NS070340/NS/NINDS NIH HHS/United States
- R01NS067305/NS/NINDS NIH HHS/United States
- 068545/Z/02/WT_/Wellcome Trust/United Kingdom
- G0700061/MRC_/Medical Research Council/United Kingdom
- 084702/Z/08/Z/WT_/Wellcome Trust/United Kingdom
- NS26799/NS/NINDS NIH HHS/United States
- G0000934/MRC_/Medical Research Council/United Kingdom
- UL1 RR024975/RR/NCRR NIH HHS/United States
- R01 NS032830/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

