Cooperation between the Hepatitis C Virus p7 and NS5B Proteins Enhances Virion Infectivity
- PMID: 26355084
- PMCID: PMC4645646
- DOI: 10.1128/JVI.01185-15
Cooperation between the Hepatitis C Virus p7 and NS5B Proteins Enhances Virion Infectivity
Abstract
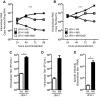
The molecular mechanisms that govern hepatitis C virus (HCV) assembly, release, and infectivity are still not yet fully understood. In the present study, we sequenced a genotype 2A strain of HCV (JFH-1) that had been cell culture adapted in Huh-7.5 cells to produce nearly 100-fold-higher viral titers than the parental strain. Sequence analysis identified nine mutations in the genome, present within both the structural and nonstructural genes. The infectious clone of this virus containing all nine culture-adapted mutations had 10-fold-higher levels of RNA replication and RNA release into the supernatant but had nearly 1,000-fold-higher viral titers, resulting in an increased specific infectivity compared to wild-type JFH-1. Two mutations, identified in the p7 polypeptide and NS5B RNA-dependent RNA polymerase, were sufficient to increase the specific infectivity of JFH-1. We found that the culture-adapted mutation in p7 promoted an increase in the size of cellular lipid droplets following transfection of viral RNA. In addition, we found that the culture-adaptive mutations in p7 and NS5B acted synergistically to enhance the specific viral infectivity of JFH-1 by decreasing the level of sphingomyelin in the virion. Overall, these results reveal a genetic interaction between p7 and NS5B that contributes to virion specific infectivity. Furthermore, our results demonstrate a novel role for the RNA-dependent RNA polymerase NS5B in HCV assembly.
Importance: Hepatitis C virus assembly and release depend on viral interactions with host lipid metabolic pathways. Here, we demonstrate that the viral p7 and NS5B proteins cooperate to promote virion infectivity by decreasing sphingomyelin content in the virion. Our data uncover a new role for the viral RNA-dependent RNA polymerase NS5B and p7 proteins in contributing to virion morphogenesis. Overall, these findings are significant because they reveal a genetic interaction between p7 and NS5B, as well as an interaction with sphingomyelin that regulates virion infectivity. Our data provide new strategies for targeting host lipid-virus interactions as potential targets for therapies against HCV infection.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures




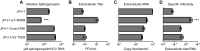
References
-
- Seeff LB. 2002. Natural history of chronic hepatitis C. Hepatology 36:S35–S46. - PubMed
-
- McHutchison JG, Lawitz EJ, Shiffman ML, Muir AJ, Galler GW, McCone J, Nyberg LM, Lee WM, Ghalib RH, Schiff ER, Galati JS, Bacon BR, Davis MN, Mukhopadhyay P, Koury K, Noviello S, Pedicone LD, Brass CA, Albrecht JK, Sulkowski MS. 2009. Peginterferon alfa-2b or alfa-2a with ribavirin for treatment of hepatitis C infection. N Engl J Med 361:580–593. doi:10.1056/NEJMoa0808010. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

